IJMS | Free Full-Text | Application and Challenge of 3rd Generation Sequencing for Clinical Bacterial Studies
Ultra Deep Sequencing of Listeria monocytogenes sRNA Transcriptome Revealed New Antisense RNAs | PLOS ONE
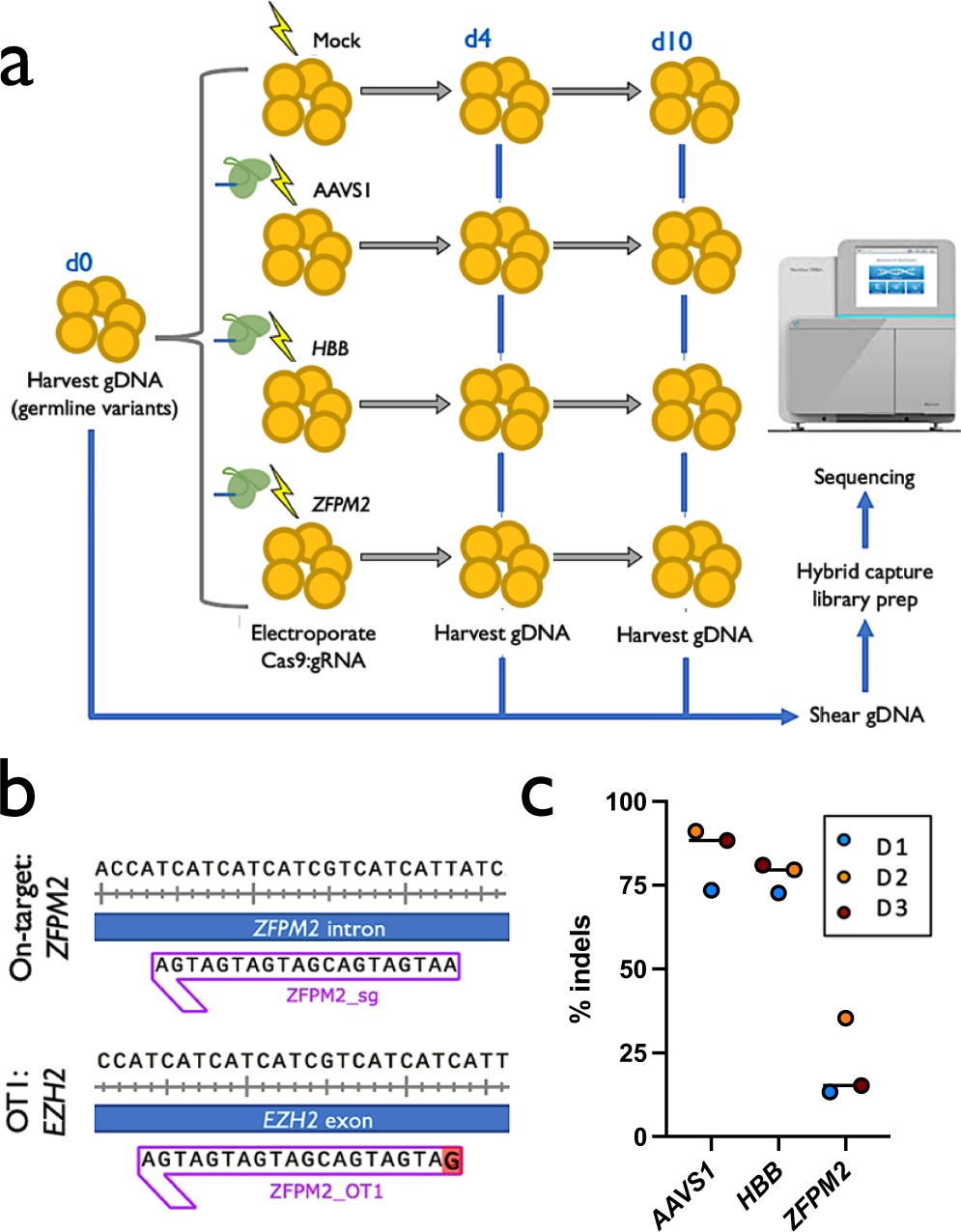
Ultra-deep sequencing validates safety of CRISPR/Cas9 genome editing in human hematopoietic stem and progenitor cells | Nature Communications

CXCR4 Mutations Detected Using Ultra-Deep Next-Generation Sequencing.... | Download Scientific Diagram
Ultra-deep massively parallel sequencing with unique molecular identifier tagging achieves comparable performance to droplet digital PCR for detection and quantification of circulating tumor DNA from lung cancer patients | PLOS ONE
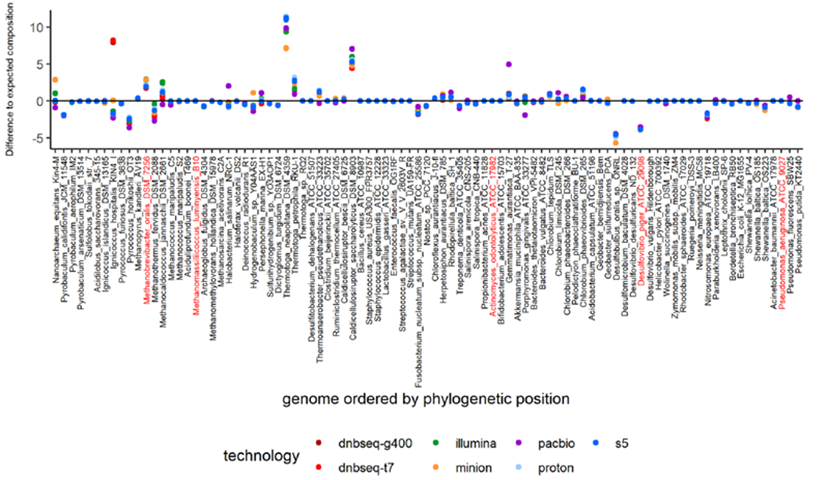
MGI's DNBSEQ-T7* Facilitates Ultra-deep Sequencing of High-complexity Metagenomic Samples-MGI-Leading Life Science Innovation

An innovative data analysis strategy for accurate next-generation sequencing detection of tumor mitochondrial DNA mutations: Molecular Therapy - Nucleic Acids
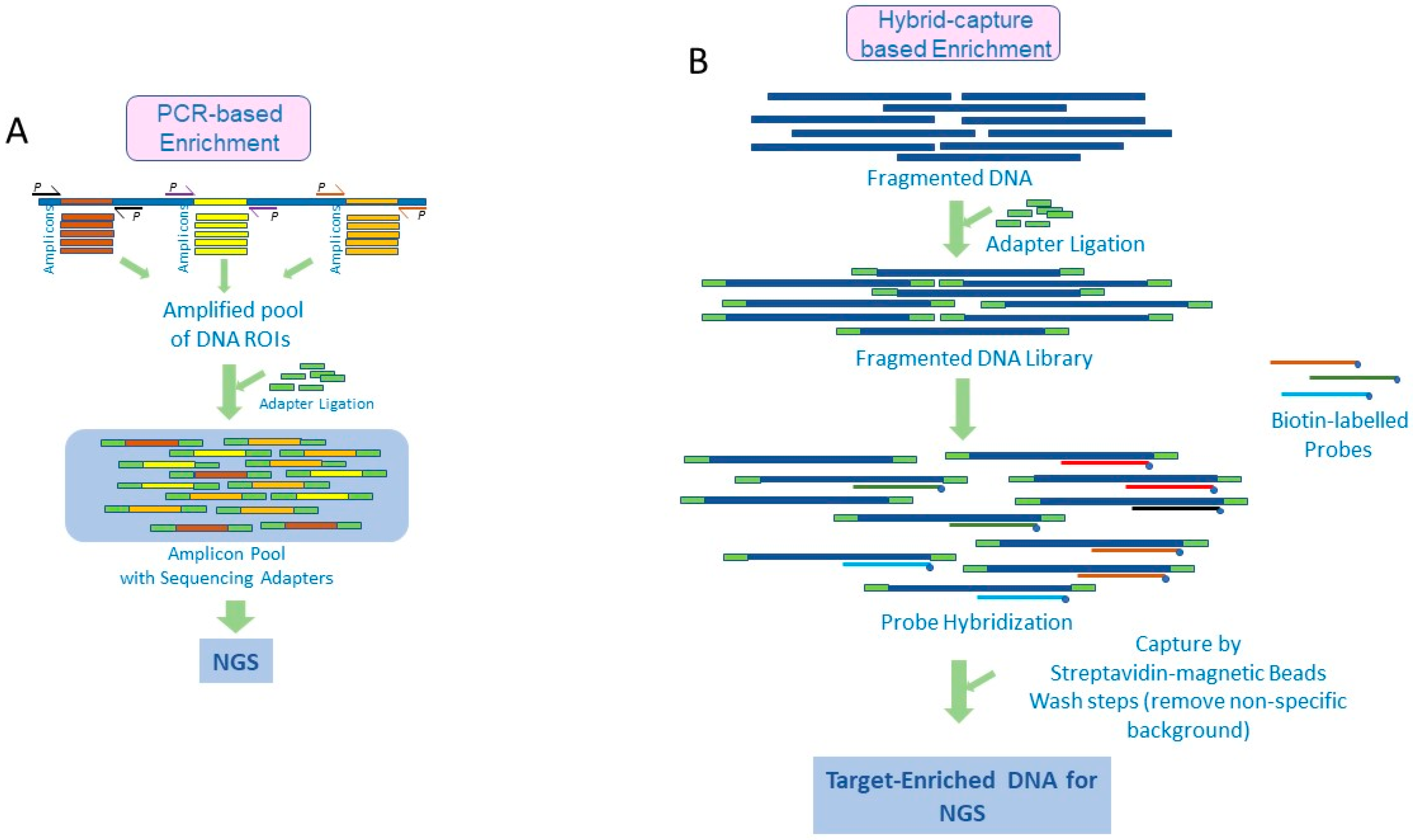
Diagnostics | Free Full-Text | Target Enrichment Approaches for Next-Generation Sequencing Applications in Oncology

Keith Slotkin on Twitter: "Please vote for my lab's entry into the @BioRender figure competition: Ultra-deep DNA methylation patterns via amplicon sequencing. By postdoc Andrea McCue. https://t.co/CiKXWX0Cug https://t.co/PywCf3EY2Y" / Twitter
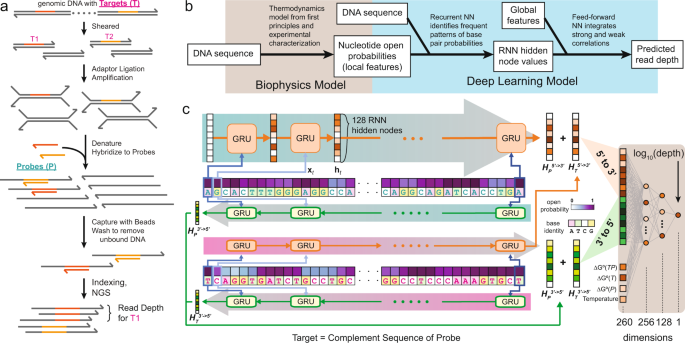
A deep learning model for predicting next-generation sequencing depth from DNA sequence | Nature Communications

Comparison of variant frequencies (%) of ultra-deep sequencing with... | Download Scientific Diagram
Ultra Deep Sequencing of Listeria monocytogenes sRNA Transcriptome Revealed New Antisense RNAs | PLOS ONE
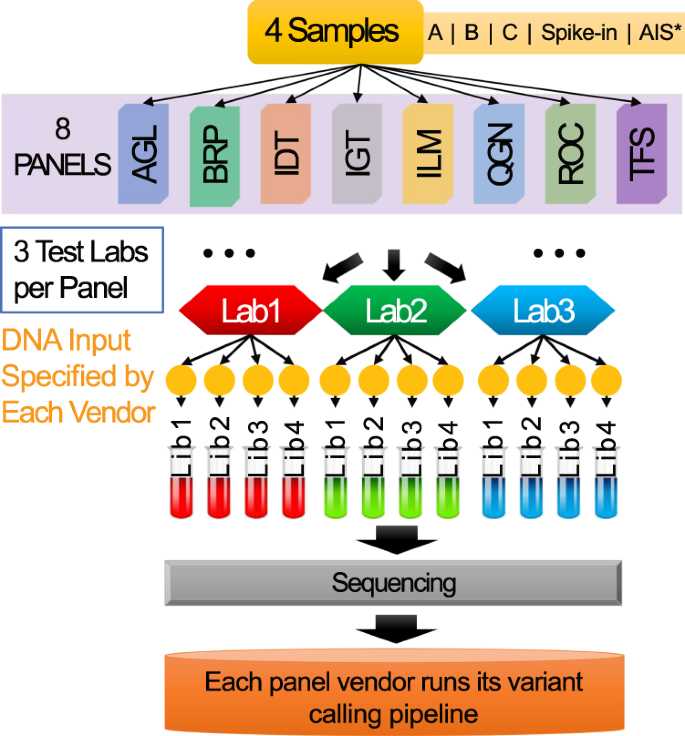
Ultra-deep multi-oncopanel sequencing of benchmarking samples with a wide range of variant allele frequencies | Scientific Data

Ultra-deep sequencing of foraminiferal microbarcodes unveils hidden richness of early monothalamous lineages in deep-sea sediments | PNAS

![PDF] Ultra-deep sequencing for the analysis of viral populations. | Semantic Scholar PDF] Ultra-deep sequencing for the analysis of viral populations. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/e953ed9fde5a4fbe86227d32bd9c09a7a2df75ed/2-Figure1-1.png)