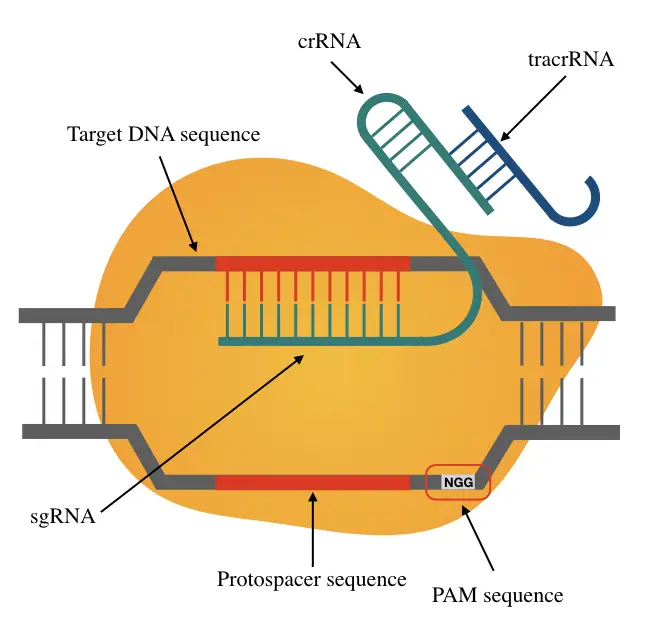
Bio-informatic analysis of CRISPR protospacer adjacent motifs (PAMs) in T4 genome | BMC Genomic Data | Full Text

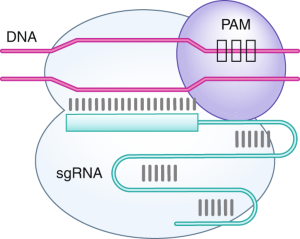
Quantification of the affinities of CRISPR–Cas9 nucleases for cognate protospacer adjacent motif (PAM) sequences - Journal of Biological Chemistry
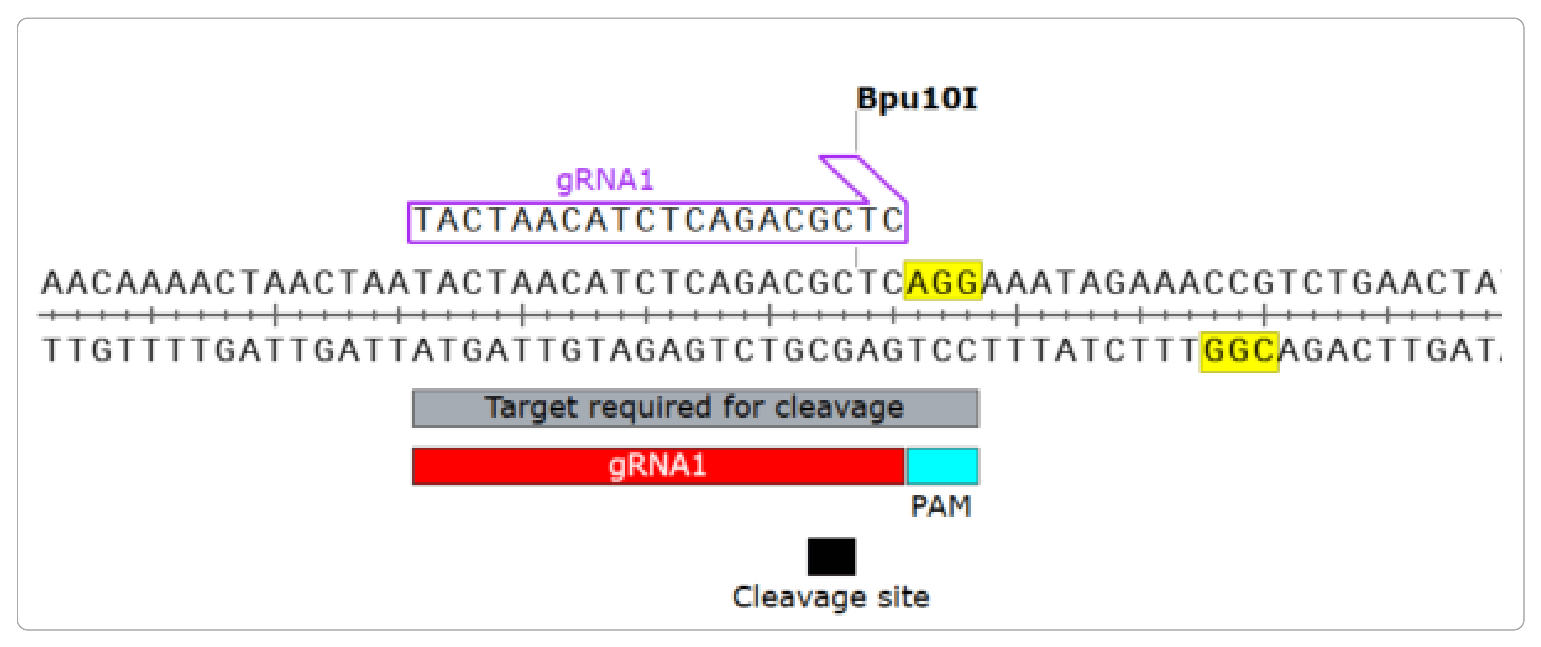
CRISPRseek: A Bioconductor Package to Identify Target-Specific Guide RNAs for CRISPR-Cas9 Genome-Editing Systems | PLOS ONE
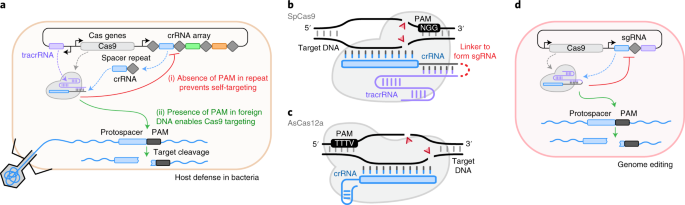
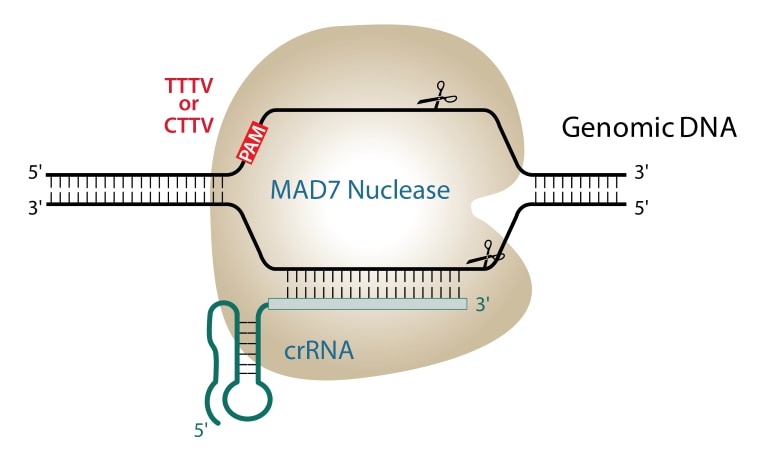
Scalable characterization of the PAM requirements of CRISPR–Cas enzymes using HT-PAMDA | Nature Protocols


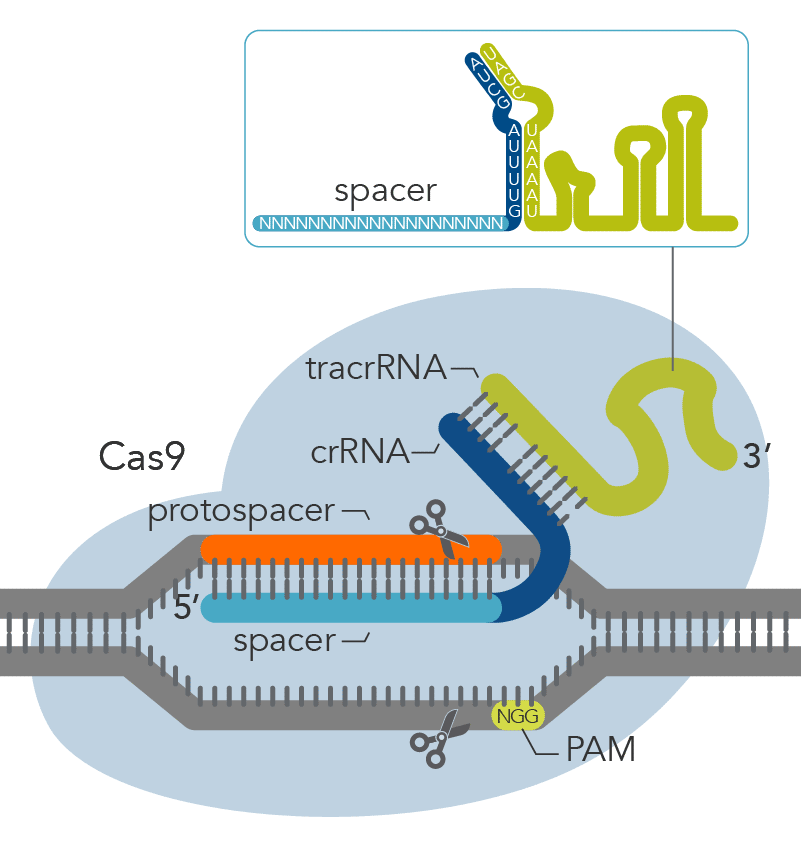

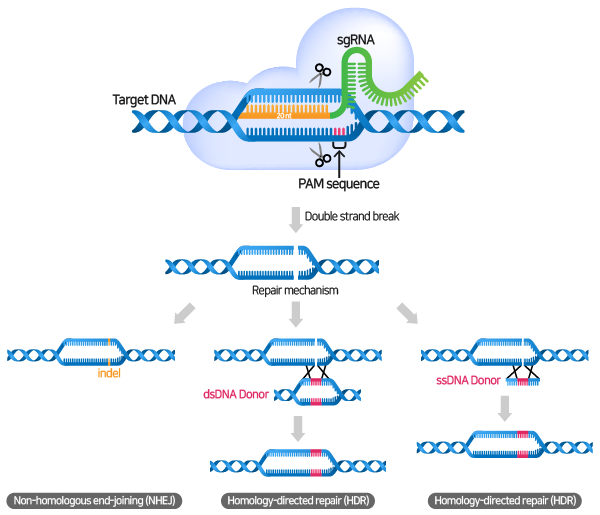
.png?sfvrsn=58c63b07_0)