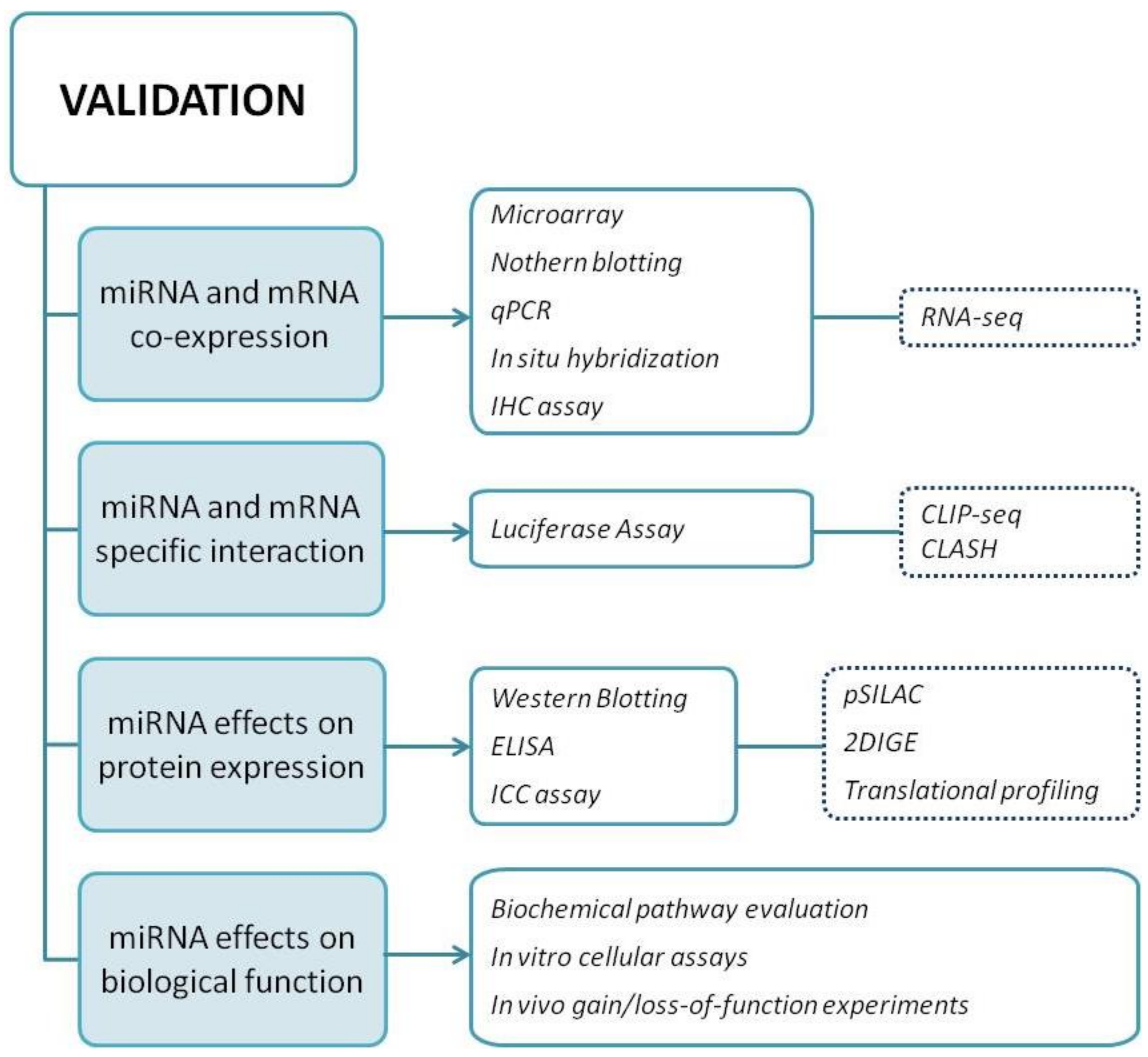
Custom target prediction with user-provided sequences. (A) Workflow of... | Download Scientific Diagram

MIPDH: A Novel Computational Model for Predicting microRNA–mRNA Interactions by DeepWalk on a Heterogeneous Network | ACS Omega

Genome wide in-silico miRNA and target network prediction from stress responsive Horsegram (Macrotyloma uniflorum) accessions | Scientific Reports

RNA-Seq workflow for miRNA–target prediction in Marchantia polymorpha.... | Download Scientific Diagram

Detection of 91 potential conserved plant microRNAs in Arabidopsis thaliana and Oryza sativa identifies important new target genes.
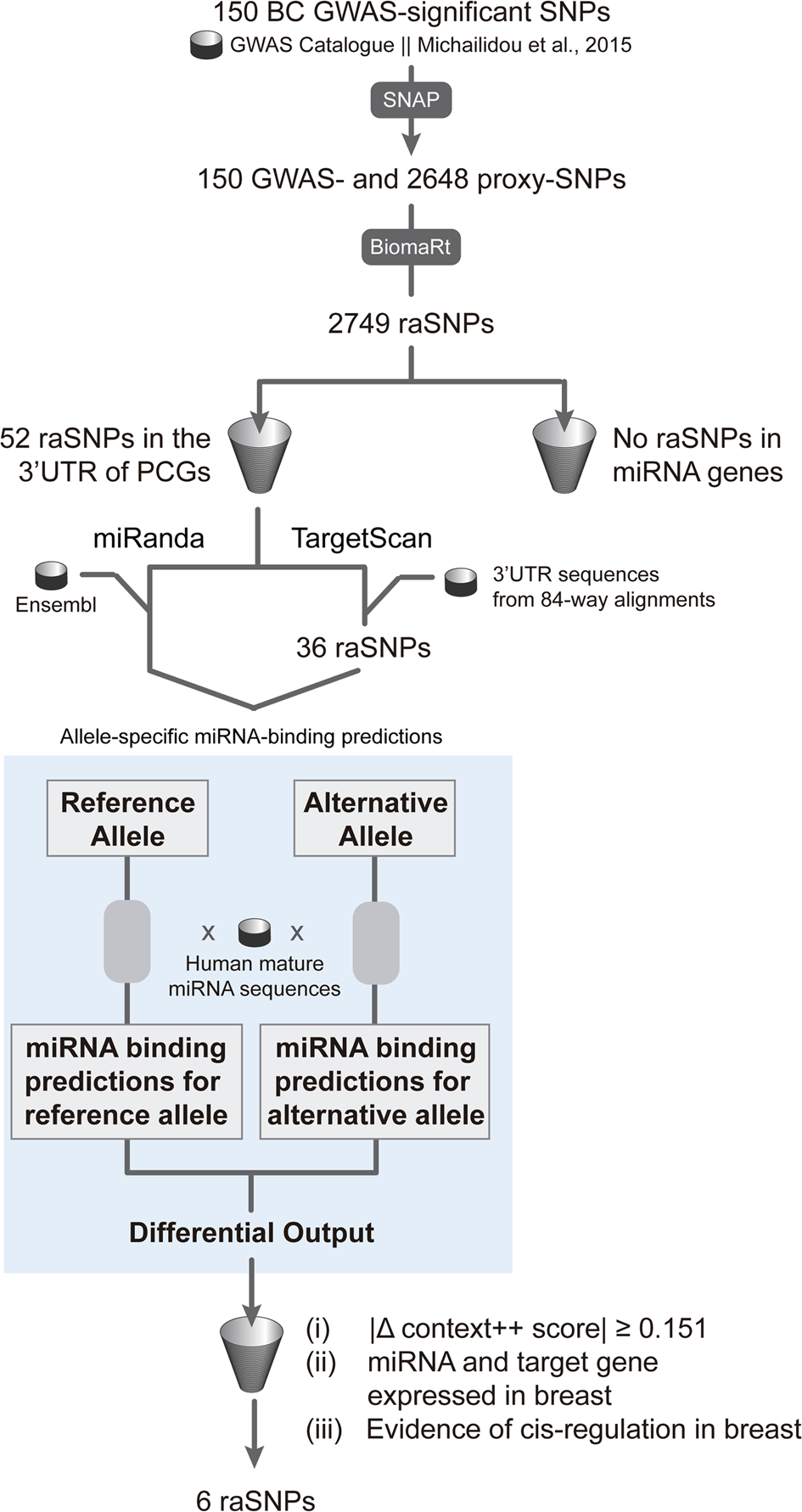
Allele-specific miRNA-binding analysis identifies candidate target genes for breast cancer risk | npj Genomic Medicine

Beyond Secondary Structure: Primary-Sequence Determinants License Pri-miRNA Hairpins for Processing: Cell
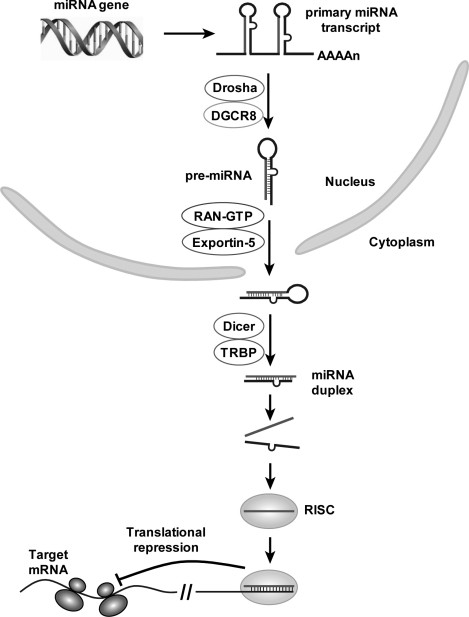
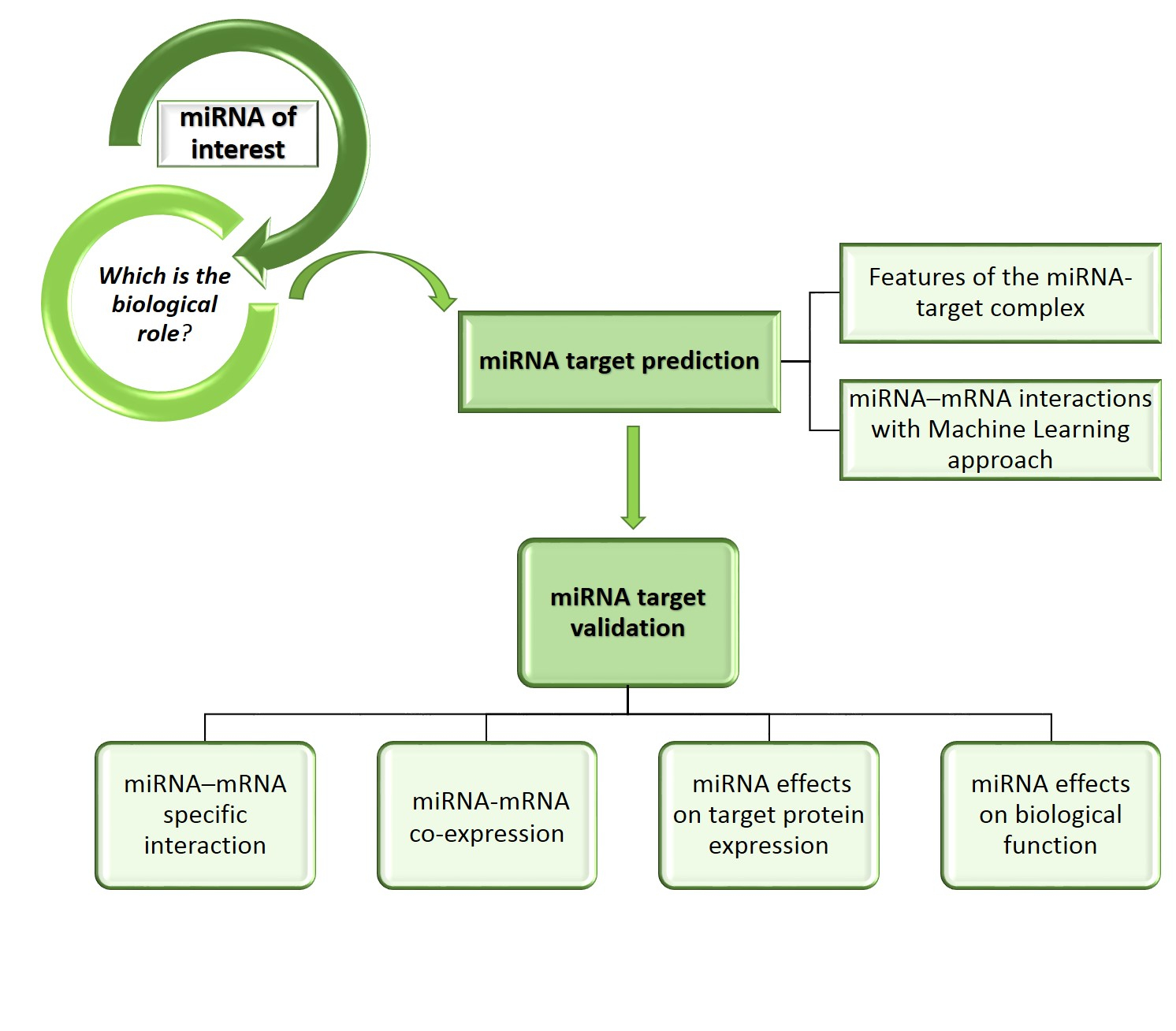
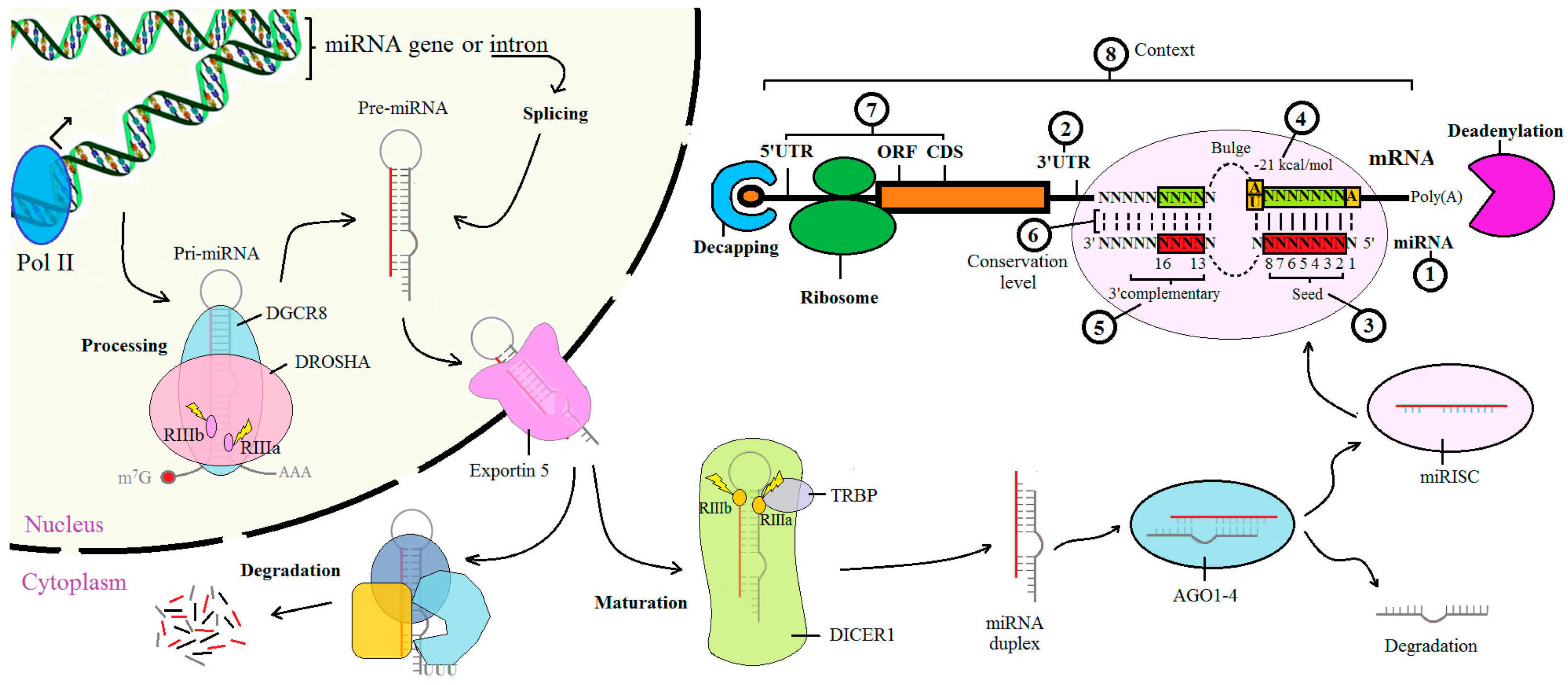
Got target?: computational methods for microRNA target prediction and their extension | Experimental & Molecular Medicine

miRWalk – Database: Prediction of possible miRNA binding sites by “walking” the genes of three genomes - ScienceDirect

MIPDH: A Novel Computational Model for Predicting microRNA–mRNA Interactions by DeepWalk on a Heterogeneous Network | ACS Omega
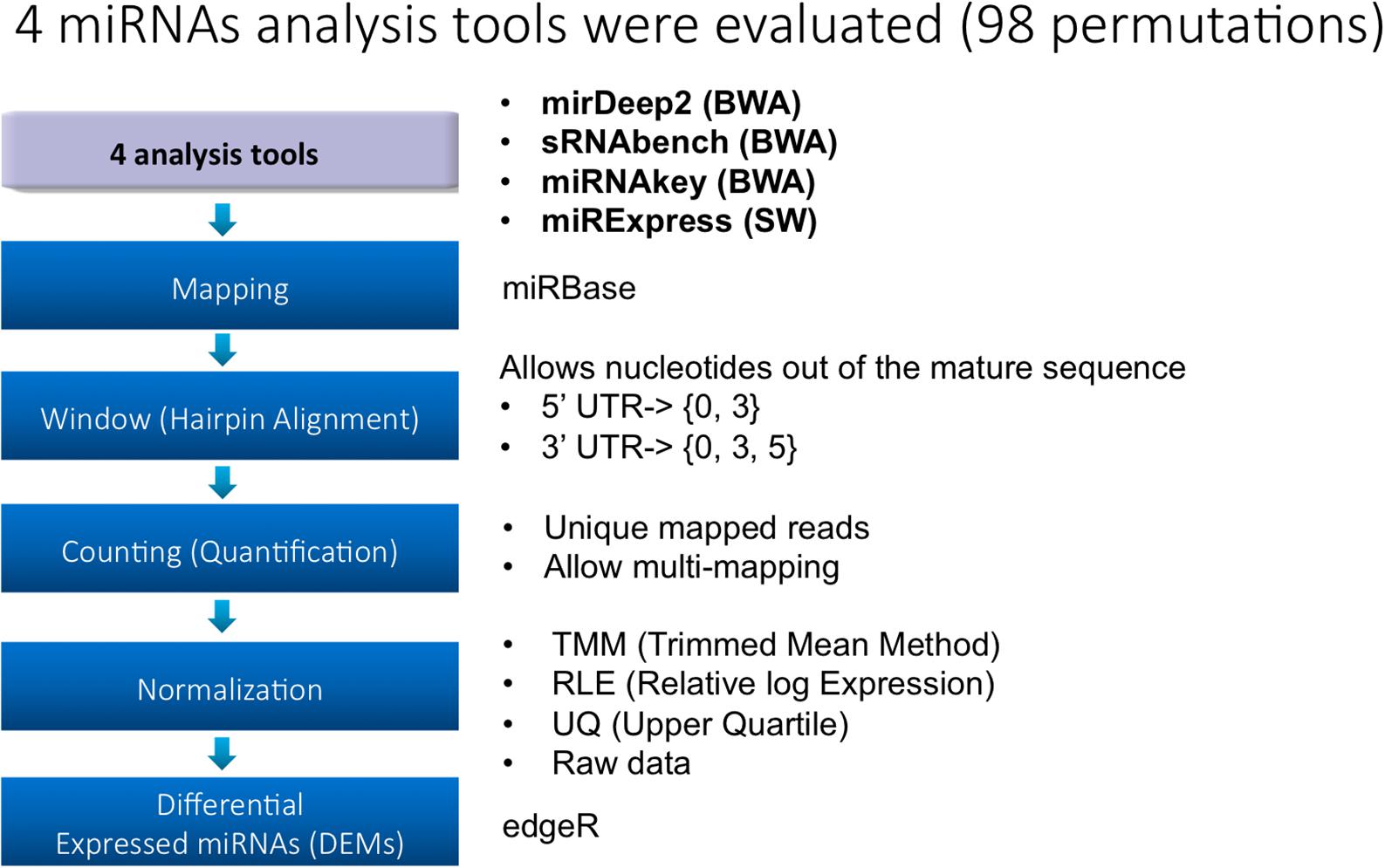

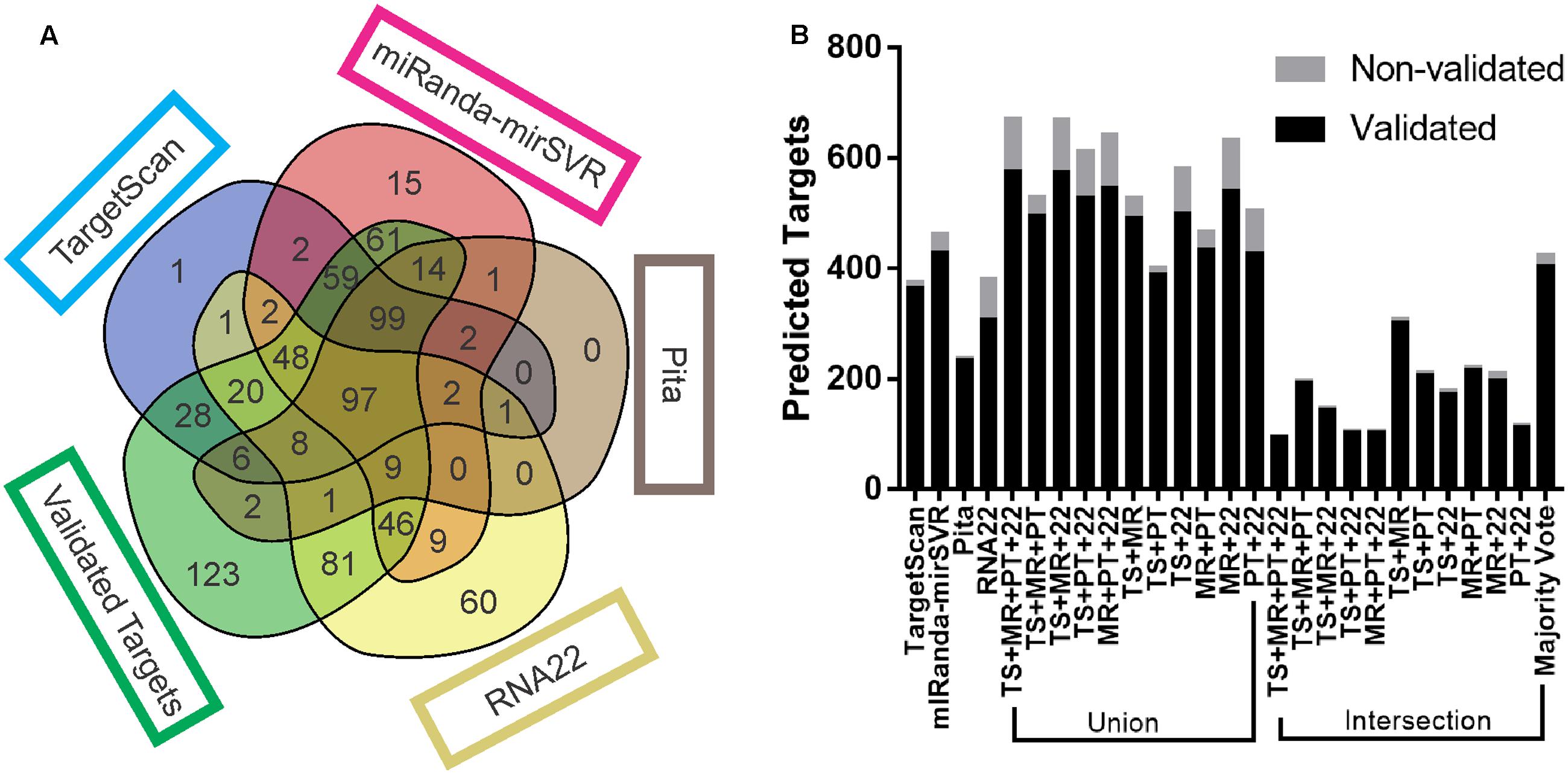
Frontiers | Evaluation of Bioinformatics Approaches for Next-Generation Sequencing Analysis of microRNAs with a Toxicogenomics Study Design



![Integrated analysis of miRNA and mRNA | 3D-Gene® [Toray DNA Chips] | TORAY Integrated analysis of miRNA and mRNA | 3D-Gene® [Toray DNA Chips] | TORAY](https://www.3d-gene.com/en/service/analysis/images/ana_004_03.gif)