Ultrahigh-Throughput Multiplexing and Sequencing of >500-Base-Pair Amplicon Regions on the Illumina HiSeq 2500 Platform | mSystems

Ultrahigh-Throughput Multiplexing and Sequencing of >500-Base-Pair Amplicon Regions on the Illumina HiSeq 2500 Platform | mSystems
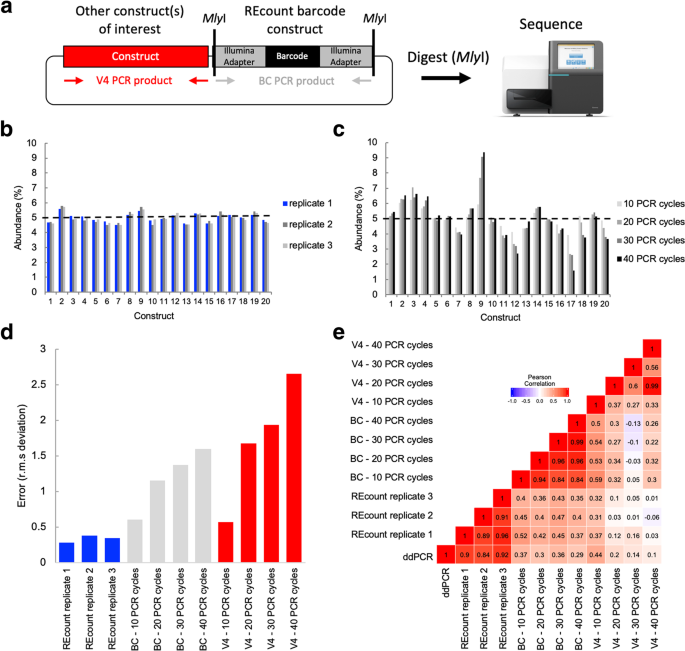
Measuring sequencer size bias using REcount: a novel method for highly accurate Illumina sequencing-based quantification | Genome Biology | Full Text
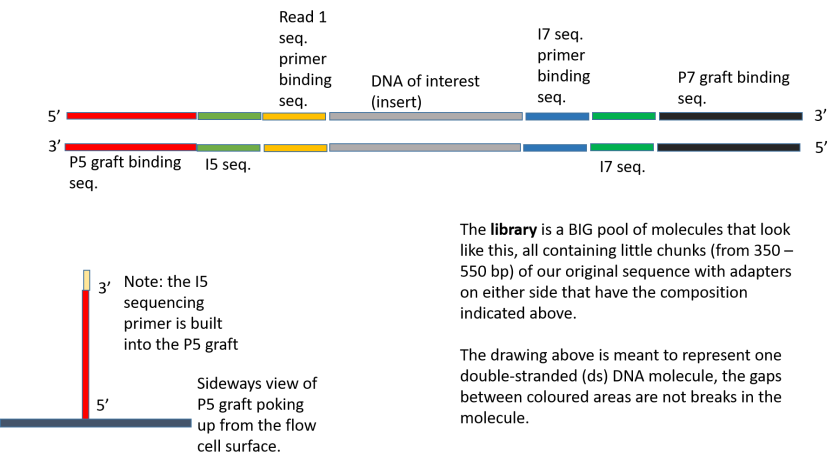
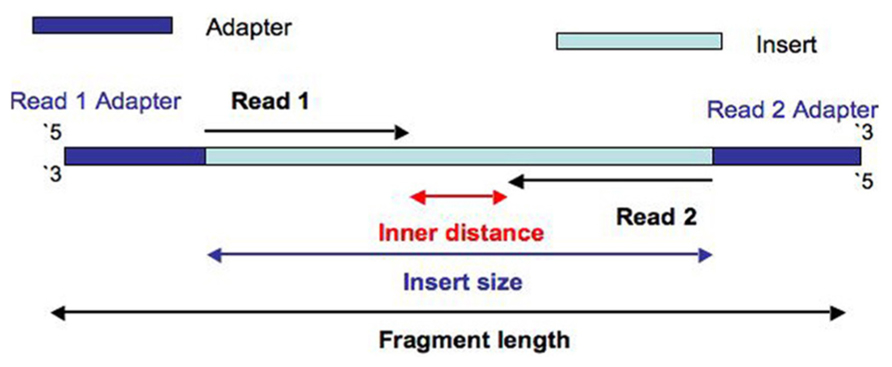
Illumina Sequencing (for Dummies) -An overview on how our samples are sequenced. – kscbioinformatics
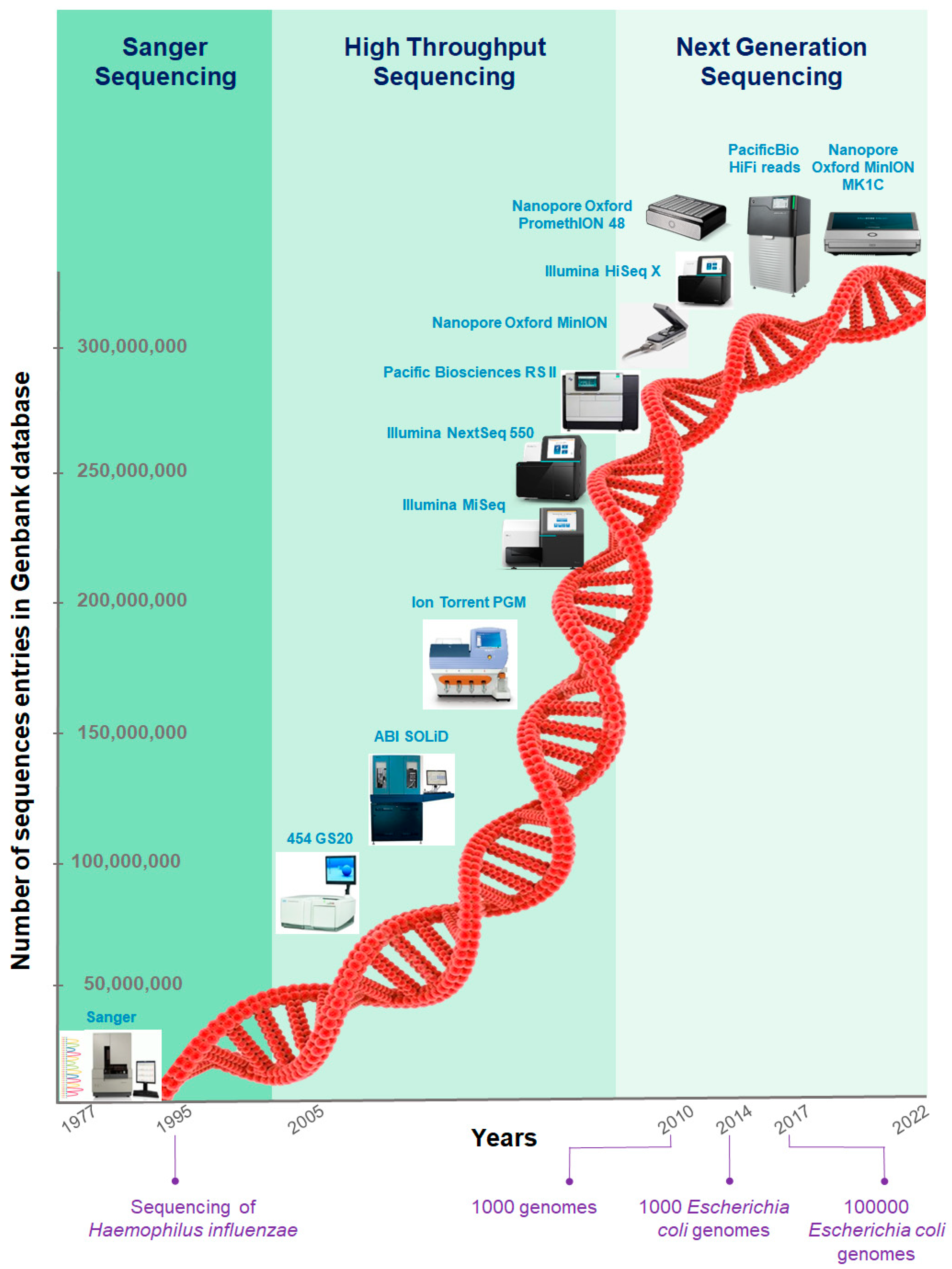
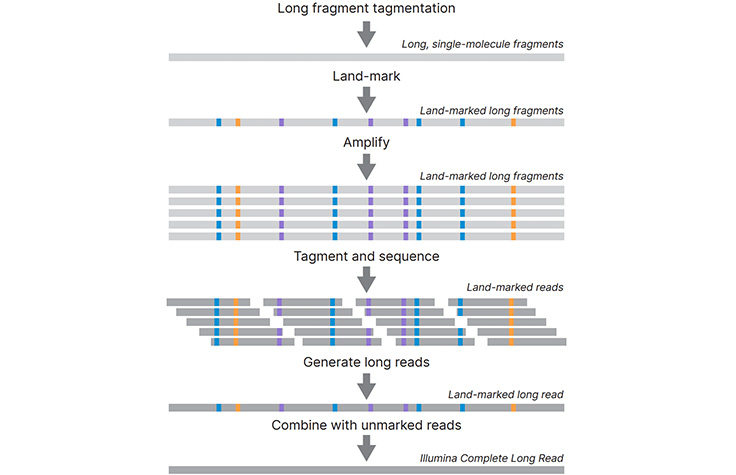
IJMS | Free Full-Text | Application and Challenge of 3rd Generation Sequencing for Clinical Bacterial Studies
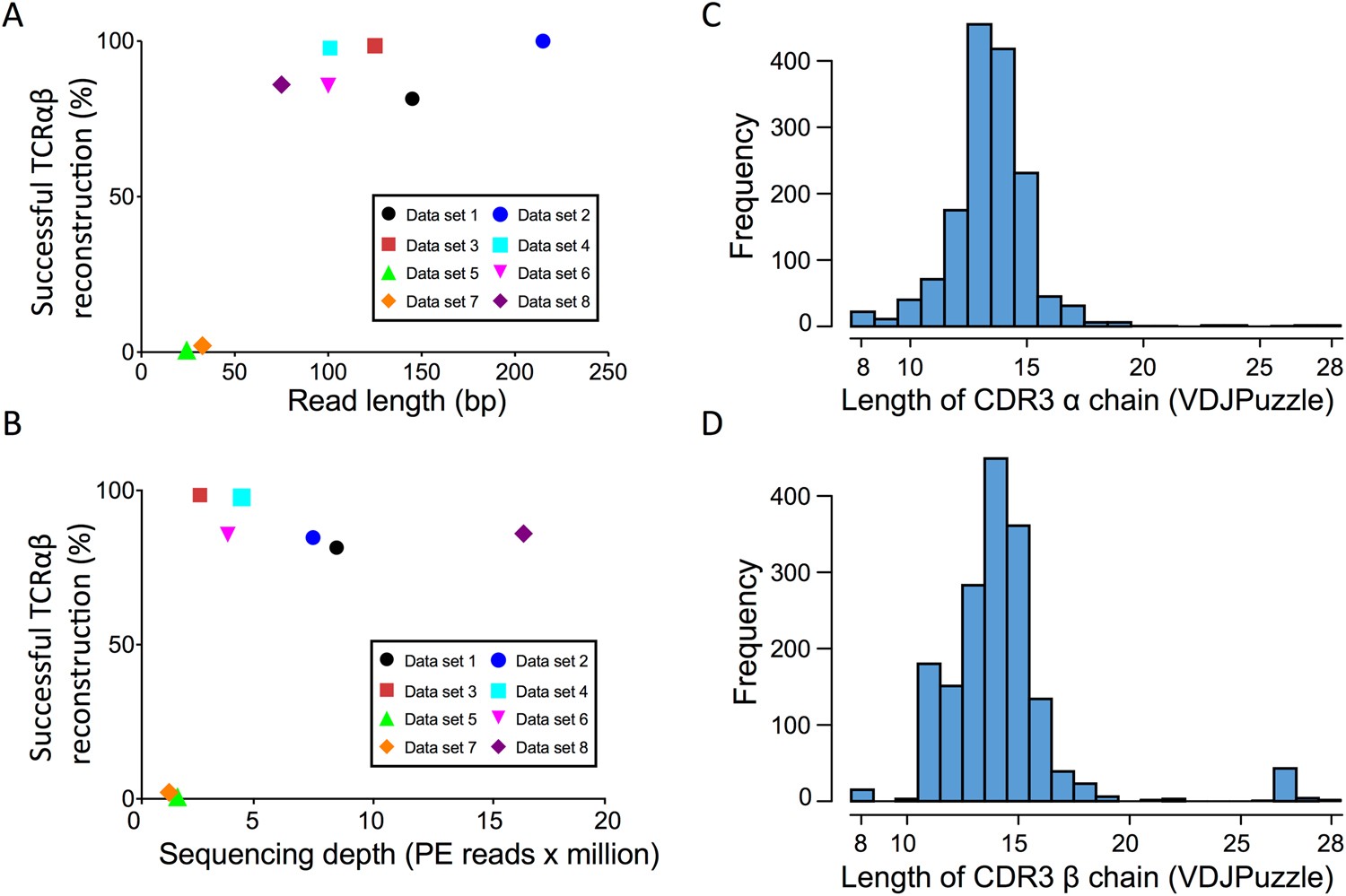
Impact of sequencing depth and read length on single cell RNA sequencing data of T cells | Scientific Reports

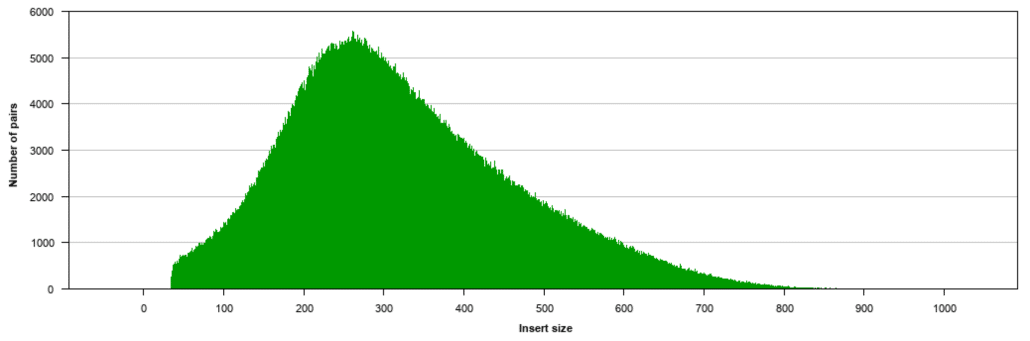
Histogram of read variants' lengths in Illumina sequencing data. The... | Download Scientific Diagram