Long walk to genomics: History and current approaches to genome sequencing and assembly - ScienceDirect

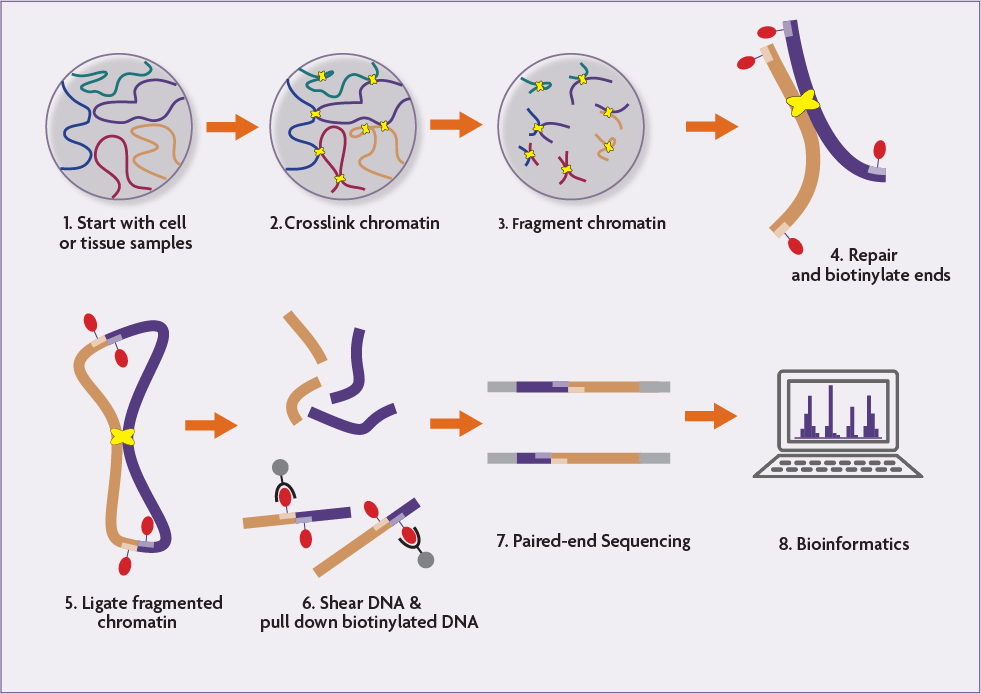
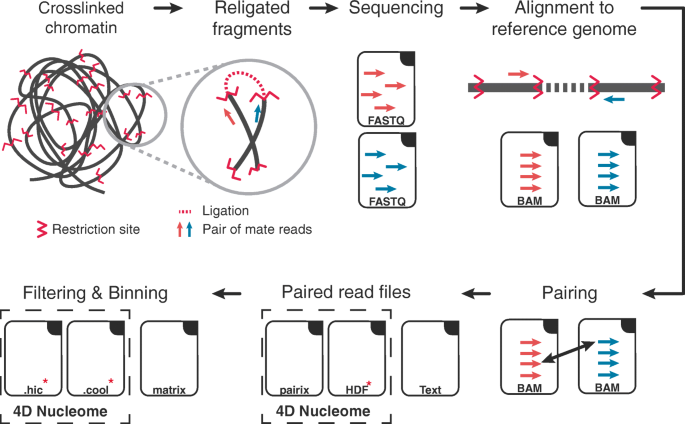
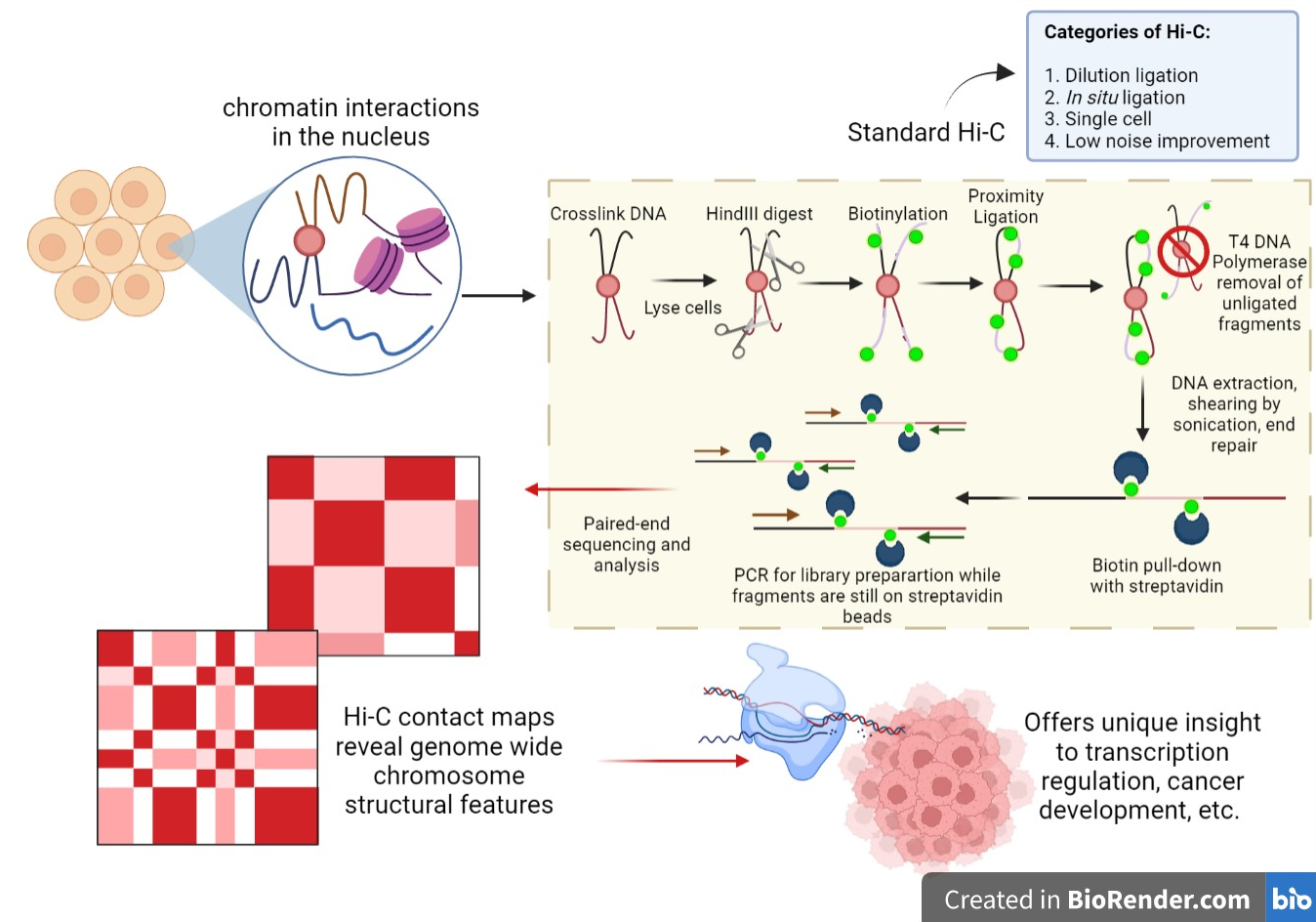
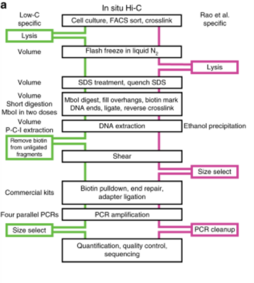
Hi‐C 3.0: Improved Protocol for Genome‐Wide Chromosome Conformation Capture - Lafontaine - 2021 - Current Protocols - Wiley Online Library
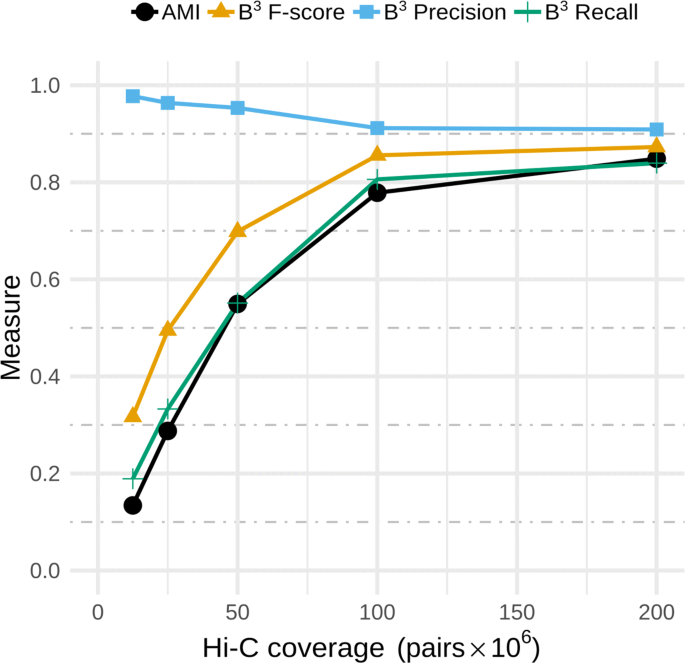
bin3C: exploiting Hi-C sequencing data to accurately resolve metagenome-assembled genomes | Genome Biology | Full Text

De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds | Science

Hi-C 2.0: An Optimized Hi-C Procedure for High-Resolution Genome-Wide Mapping of Chromosome Conformation | bioRxiv
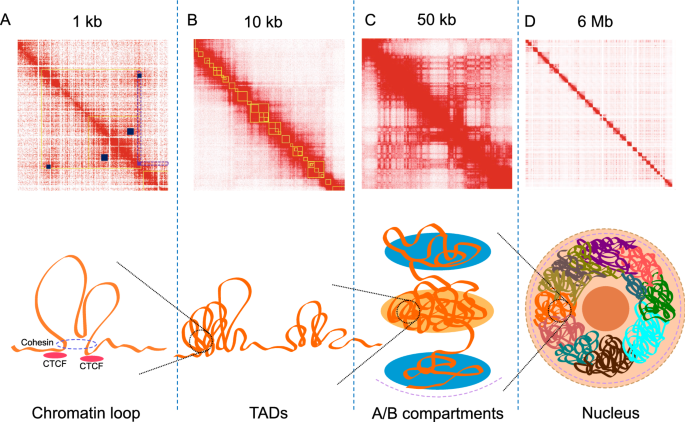
Seeing the forest through the trees: prioritising potentially functional interactions from Hi-C | Epigenetics & Chromatin | Full Text
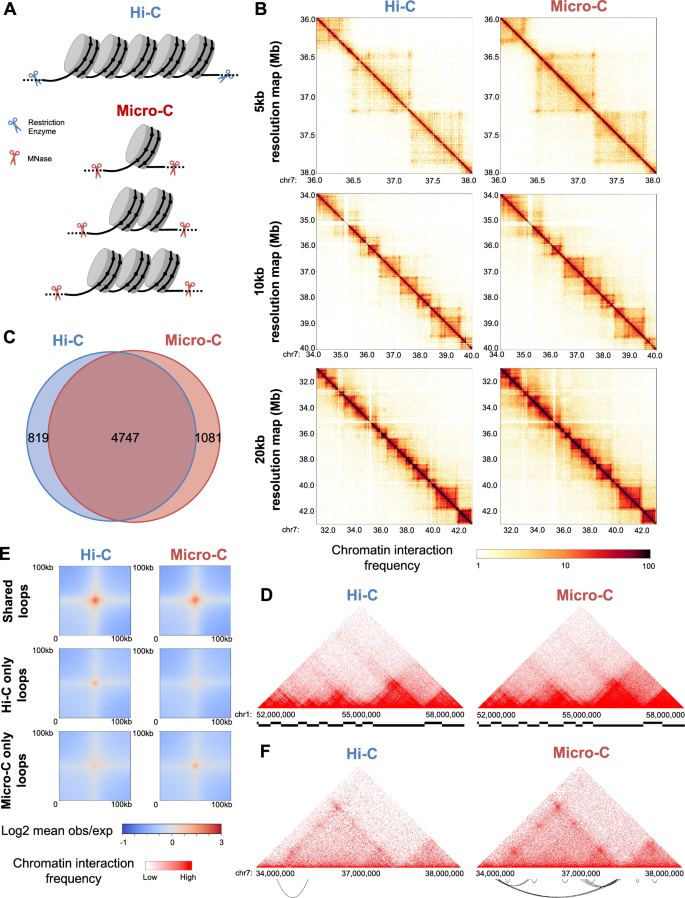
Characterizing chromatin interactions of regulatory elements and nucleosome positions, using Hi-C, Micro-C, and promoter capture Micro-C | Epigenetics & Chromatin | Full Text

BL-Hi-C is an efficient and sensitive approach for capturing structural and regulatory chromatin interactions | Nature Communications