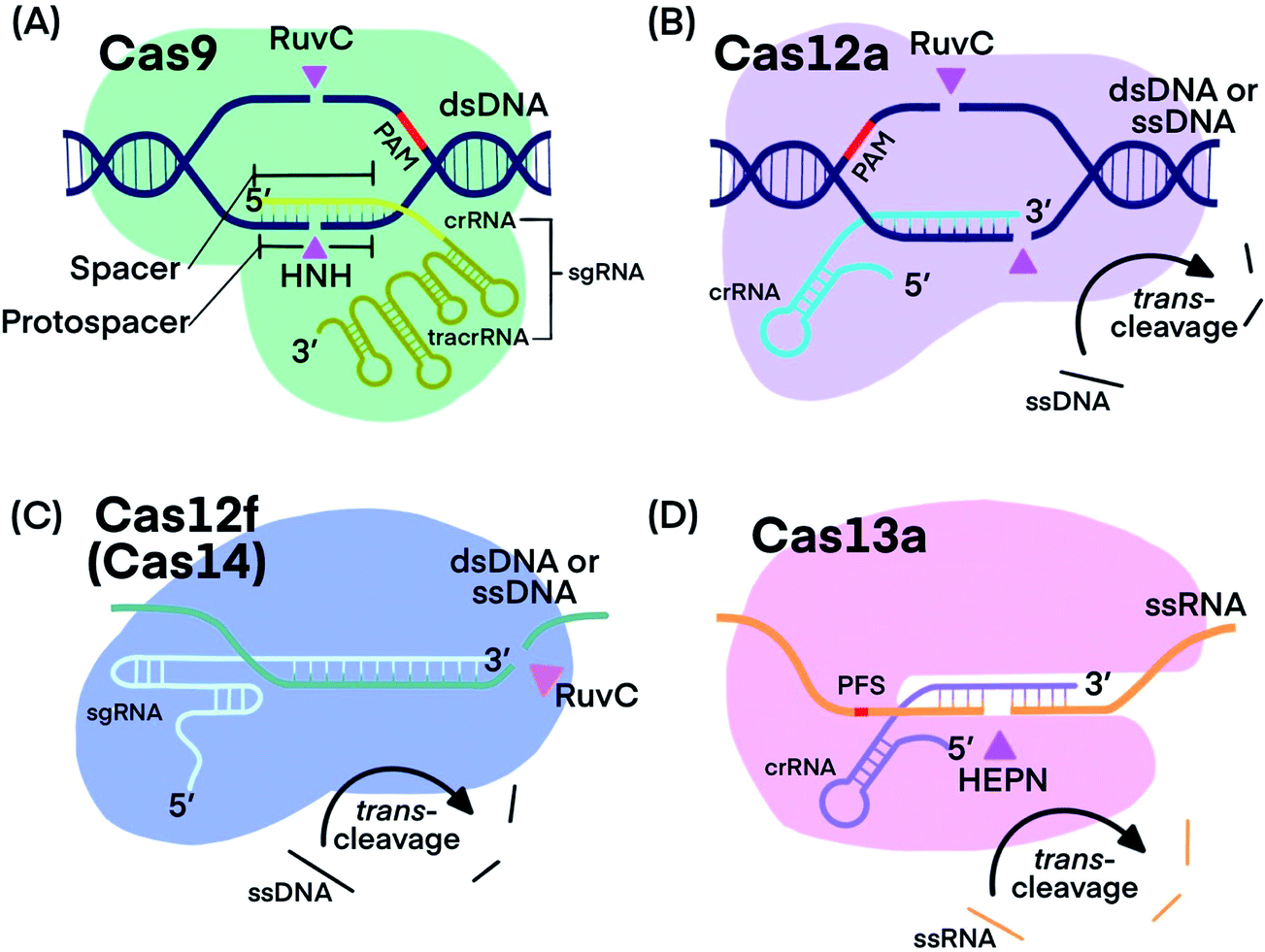
CRISPR technology incorporating amplification strategies: molecular assays for nucleic acids, proteins, and small molecules - Chemical Science (RSC Publishing) DOI:10.1039/D0SC06973F
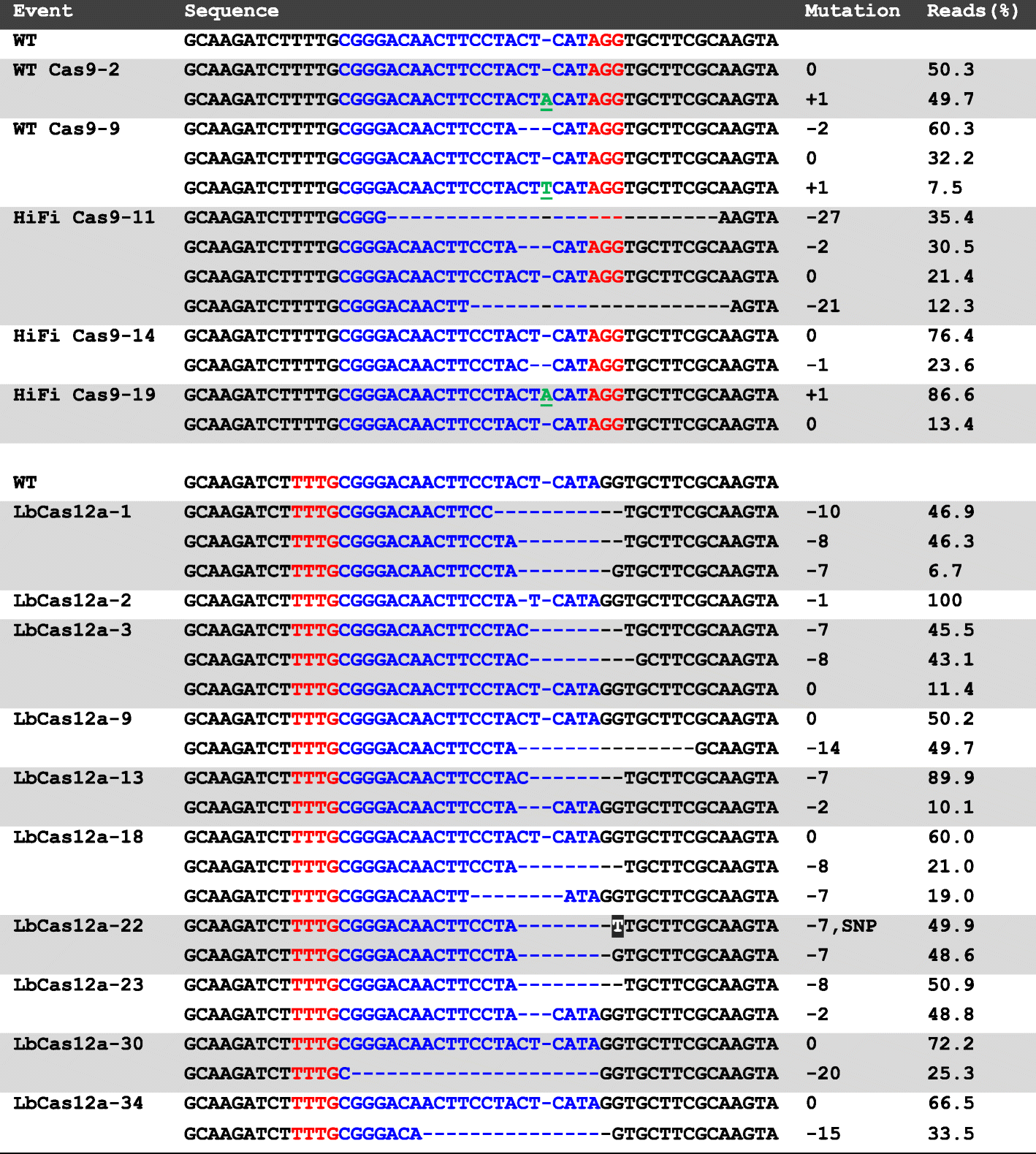
Comparison of CRISPR-Cas9/Cas12a Ribonucleoprotein Complexes for Genome Editing Efficiency in the Rice Phytoene Desaturase (OsPDS) Gene | Rice | Full Text
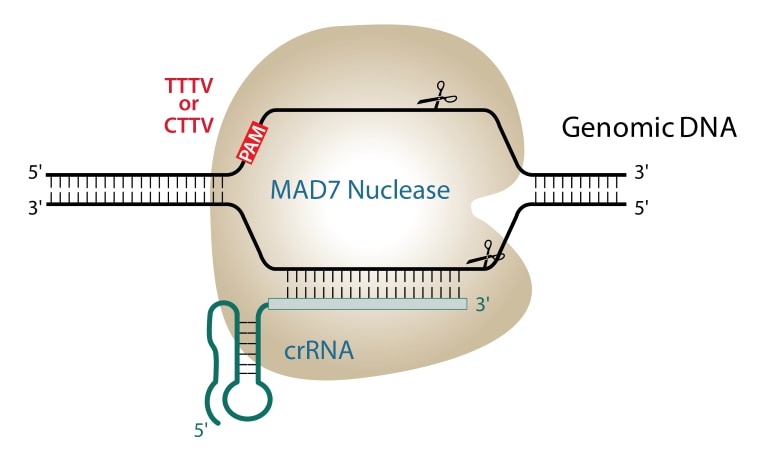
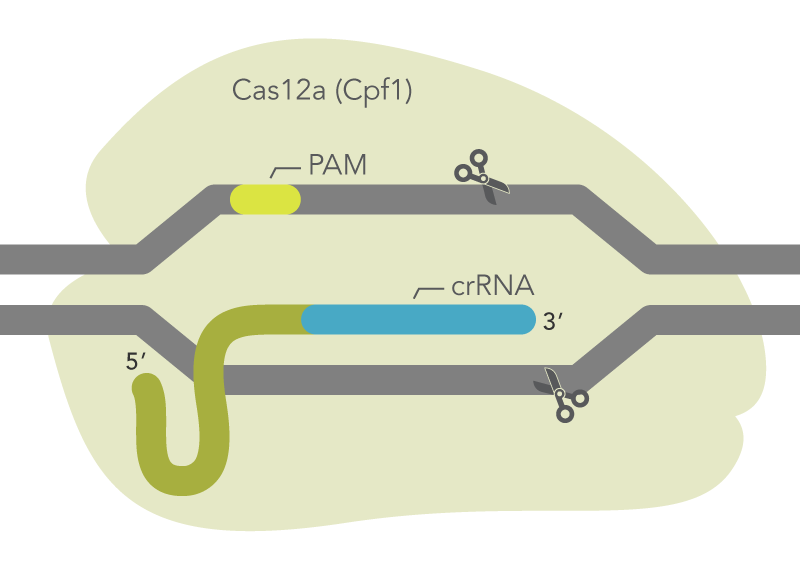
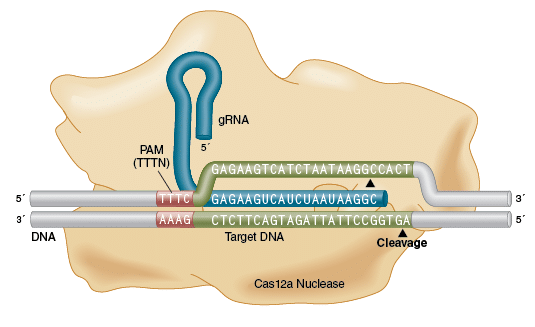
Crispr-cas12/cpf1 recognizes TTTN protospacer adjacent motif (PAM) and... | Download Scientific Diagram

A CRISPR-Cas12a-derived biosensing platform for the highly sensitive detection of diverse small molecules | Nature Communications

Scalable characterization of the PAM requirements of CRISPR–Cas enzymes using HT-PAMDA | Nature Protocols

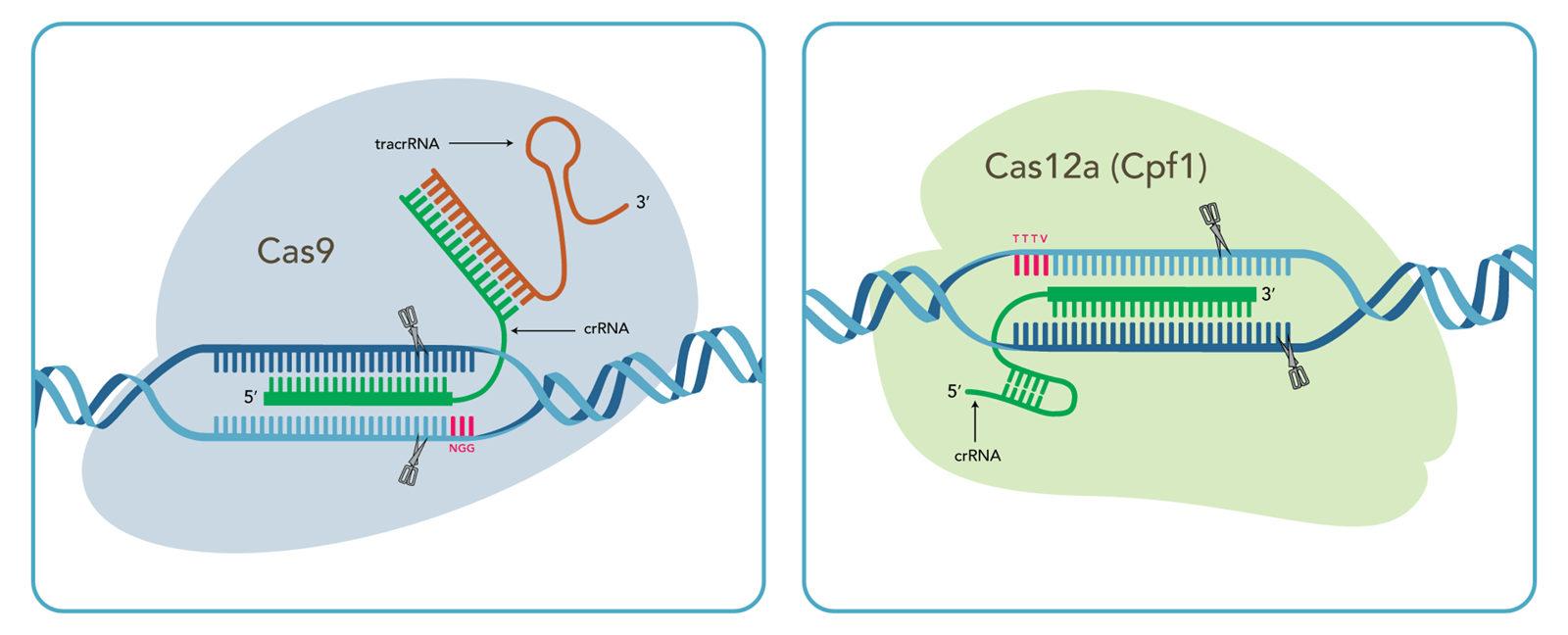
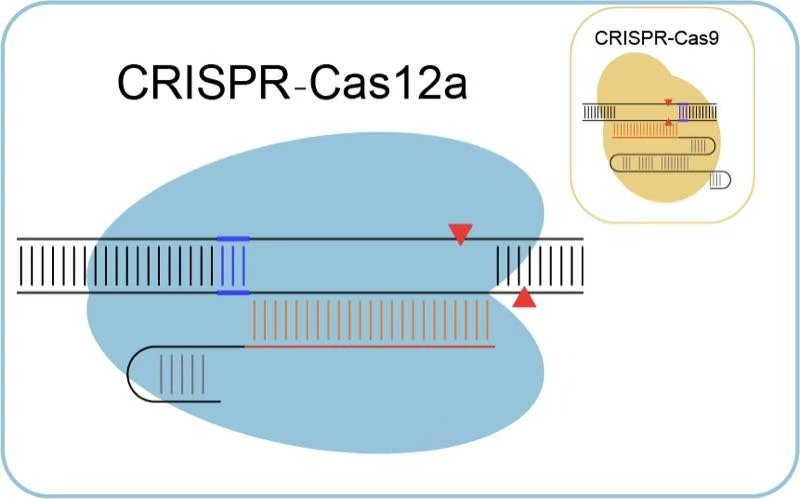
Schematic overview of CRISPR–Cas12a- and CRISPR–Cas9-mediated genome... | Download Scientific Diagram
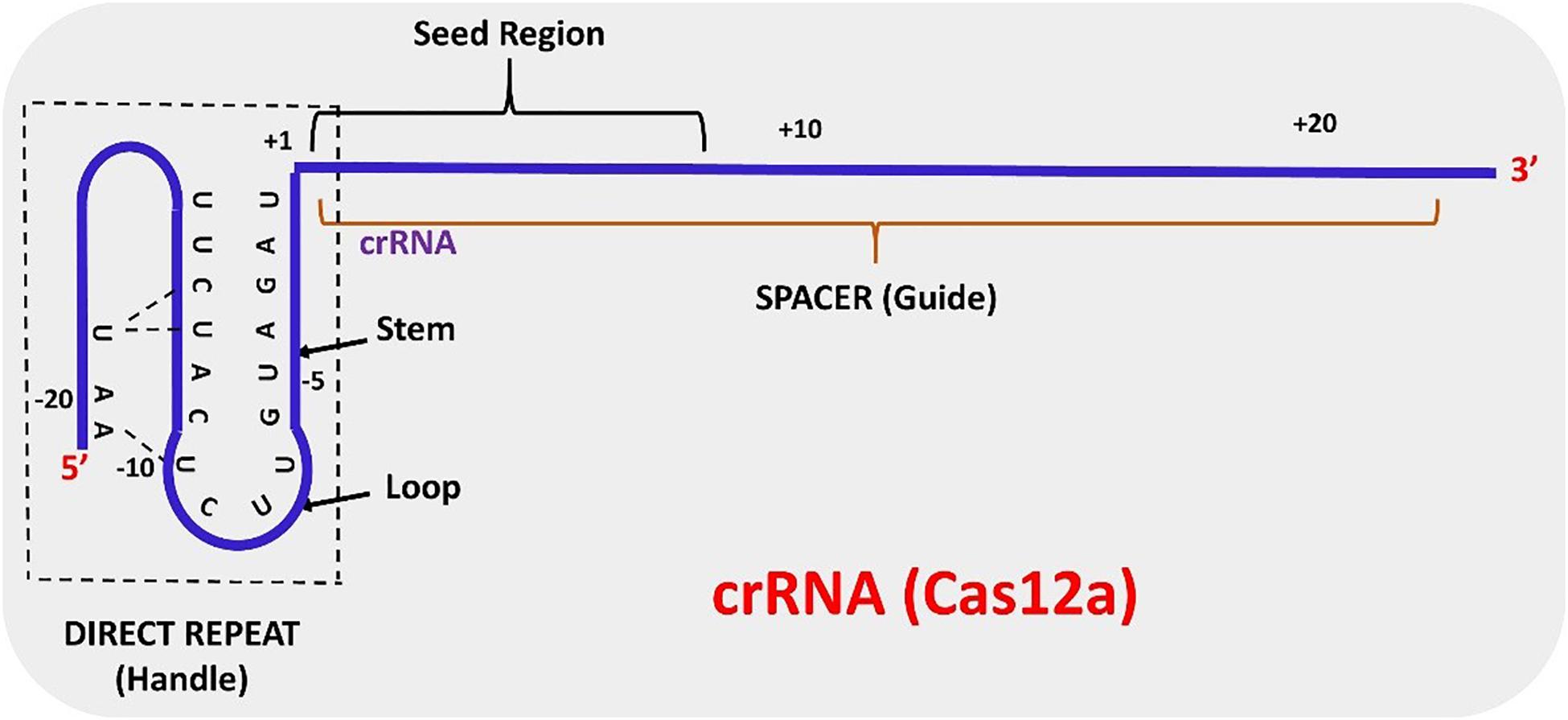
Frontiers | CRISPR-Cas12a (Cpf1): A Versatile Tool in the Plant Genome Editing Tool Box for Agricultural Advancement
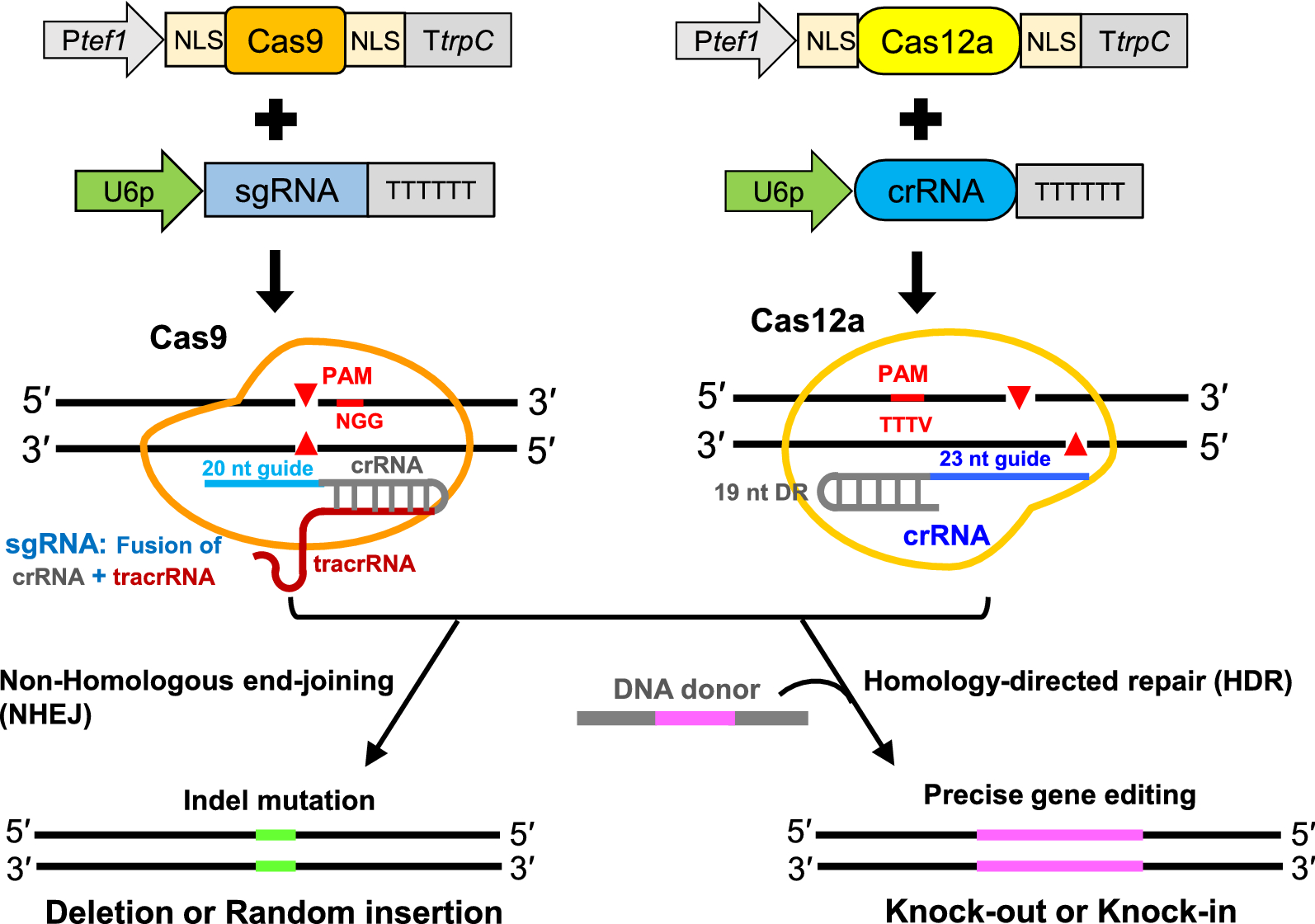
Upgrading of efficient and scalable CRISPR–Cas-mediated technology for genetic engineering in thermophilic fungus Myceliophthora thermophila | Biotechnology for Biofuels and Bioproducts | Full Text
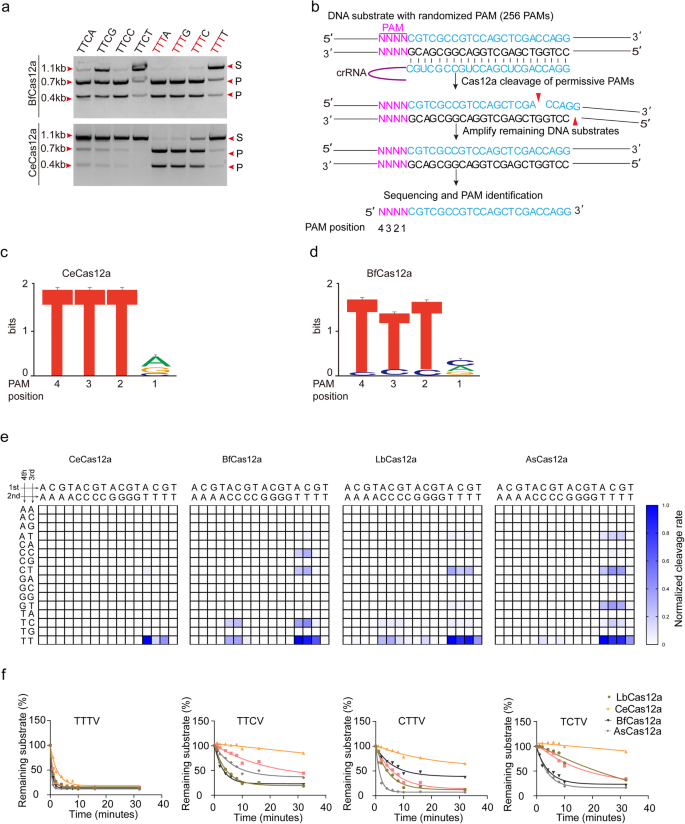
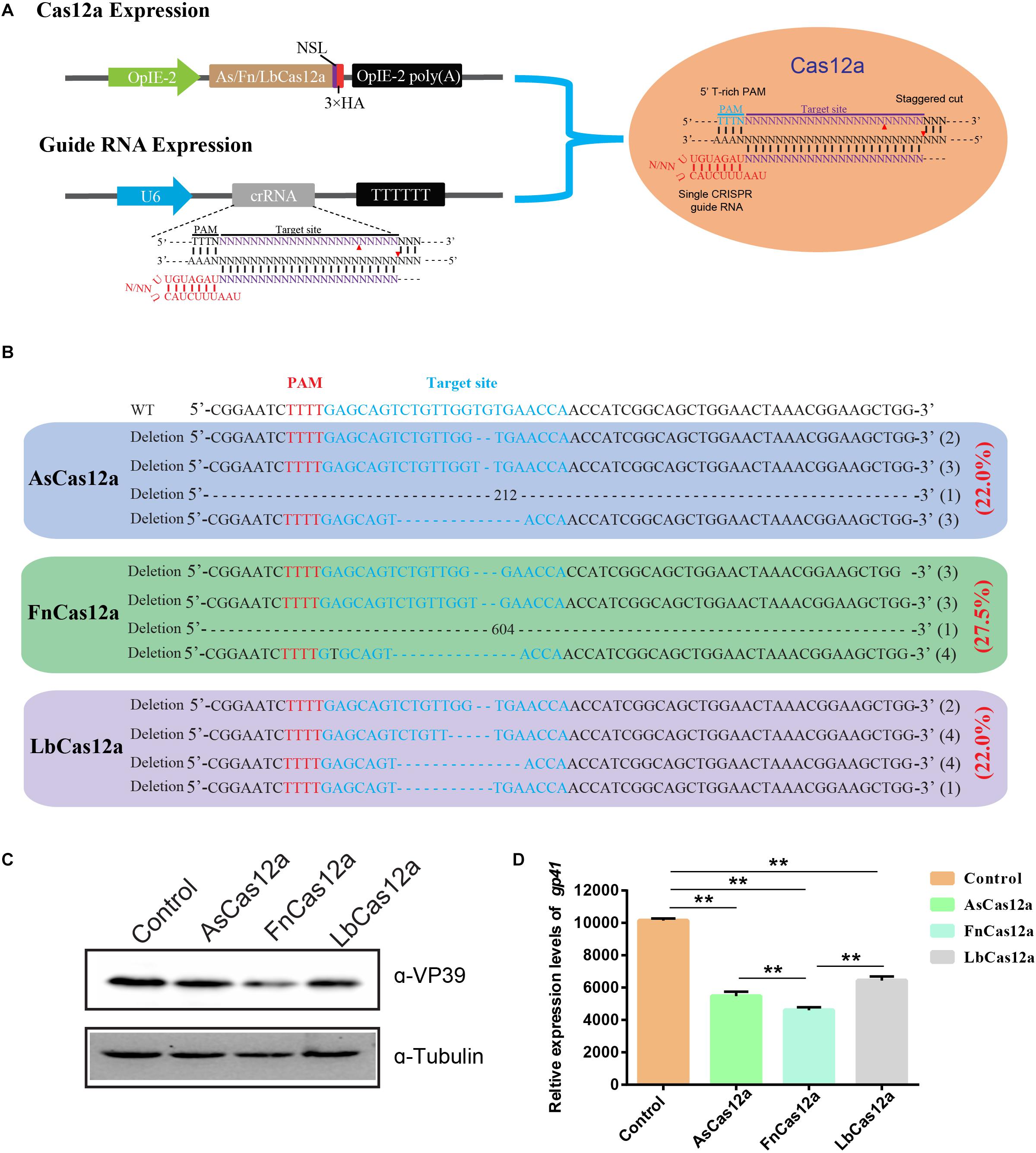
A Cas12a ortholog with stringent PAM recognition followed by low off-target editing rates for genome editing | Genome Biology | Full Text