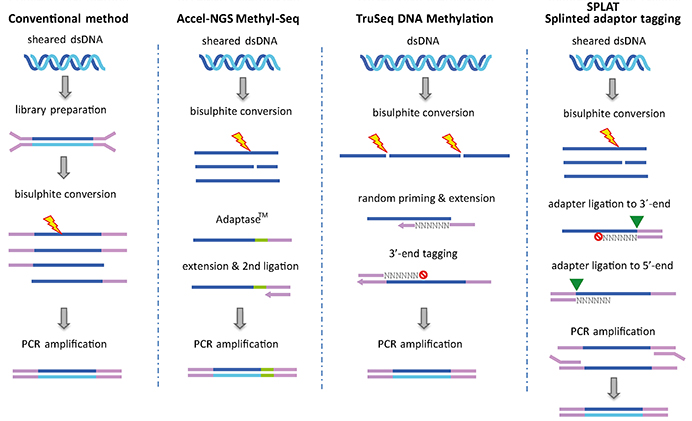
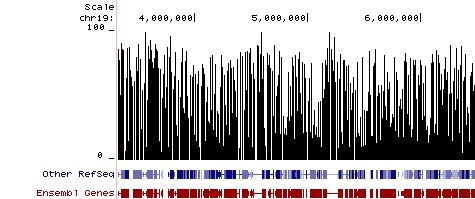
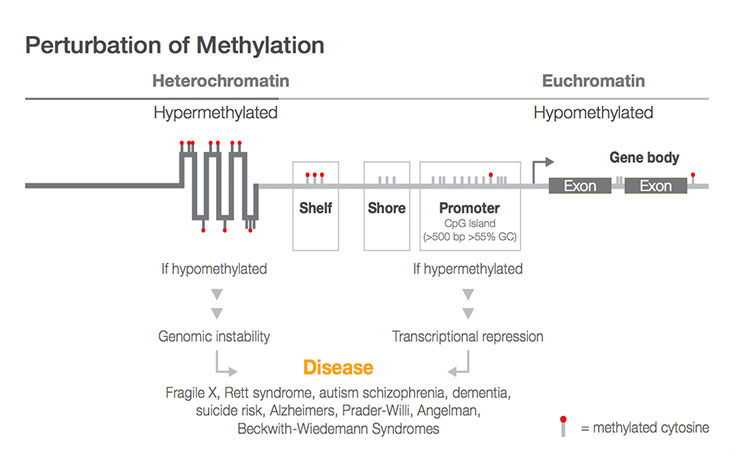
BiocMAP: A Bioconductor-friendly, GPU-Accelerated Pipeline for Bisulfite-Sequencing Data | L. Collado-Torres
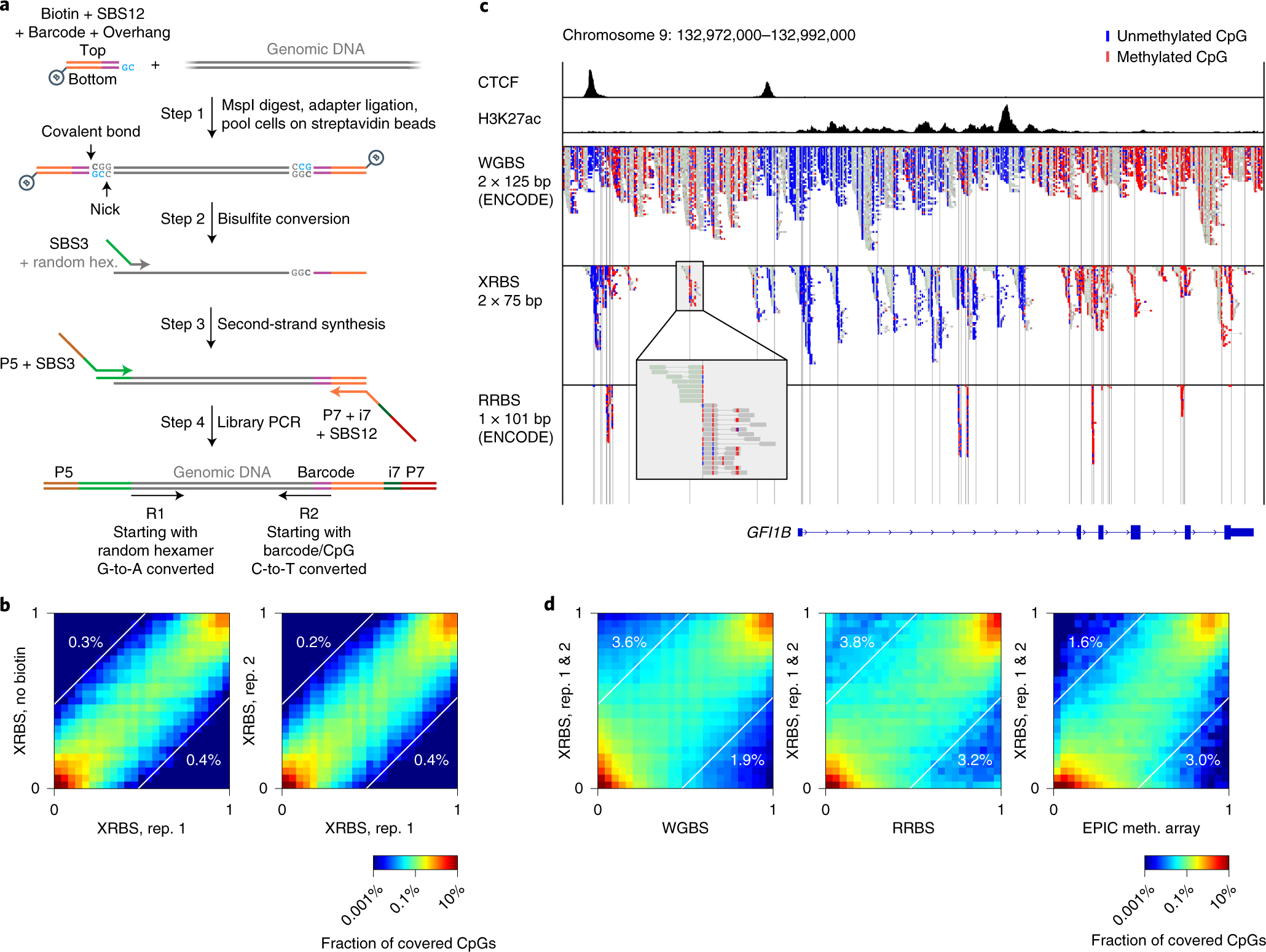
Extended-representation bisulfite sequencing of gene regulatory elements in multiplexed samples and single cells | Nature Biotechnology

MethylScore, a pipeline for accurate and context-aware identification of differentially methylated regions from population-scale plant whole-genome bisulfite sequencing data | Quantitative Plant Biology | Cambridge Core
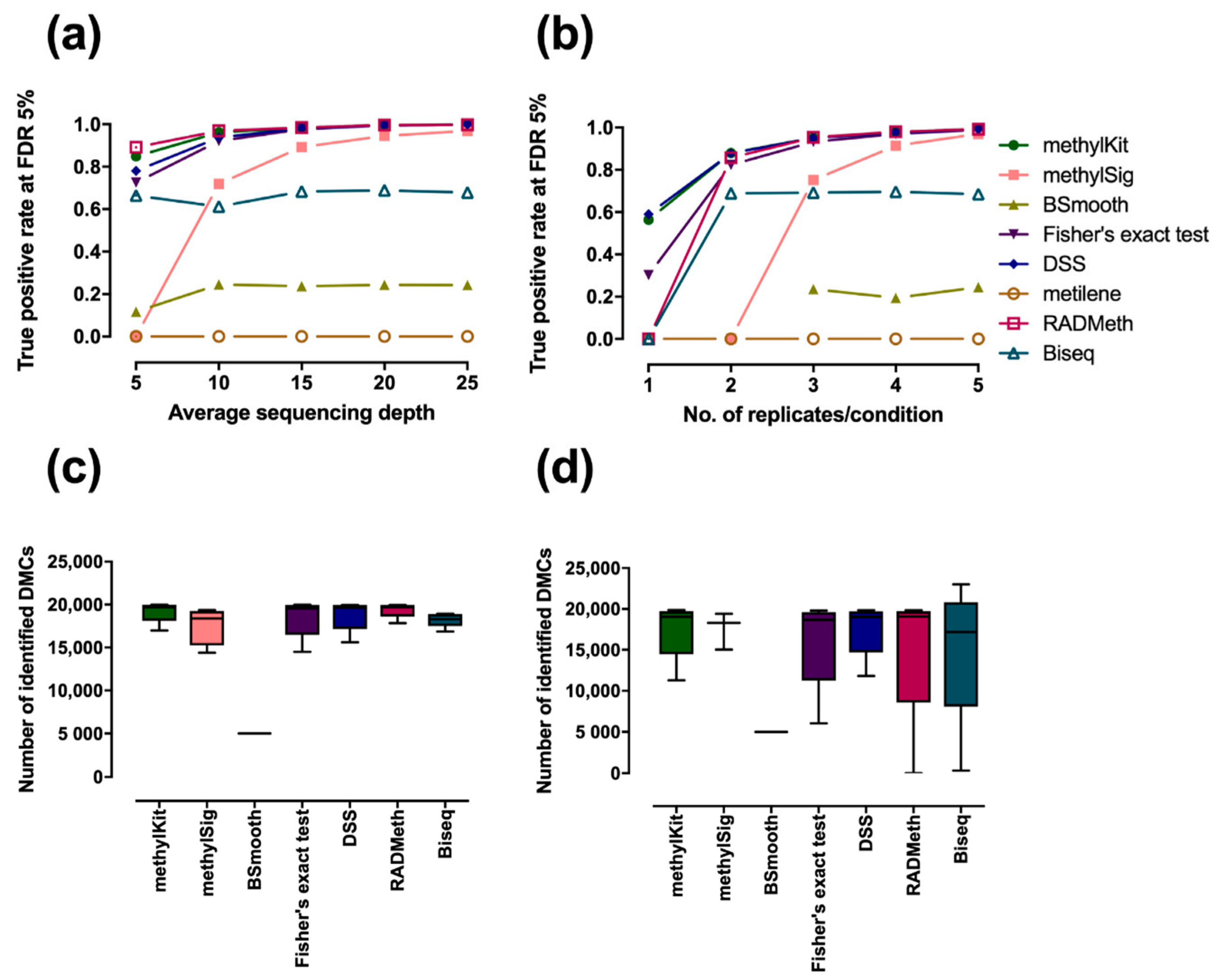
IJERPH | Free Full-Text | Comprehensive Evaluation of Differential Methylation Analysis Methods for Bisulfite Sequencing Data

Figures and data in Genome-wide detection of imprinted differentially methylated regions using nanopore sequencing | eLife
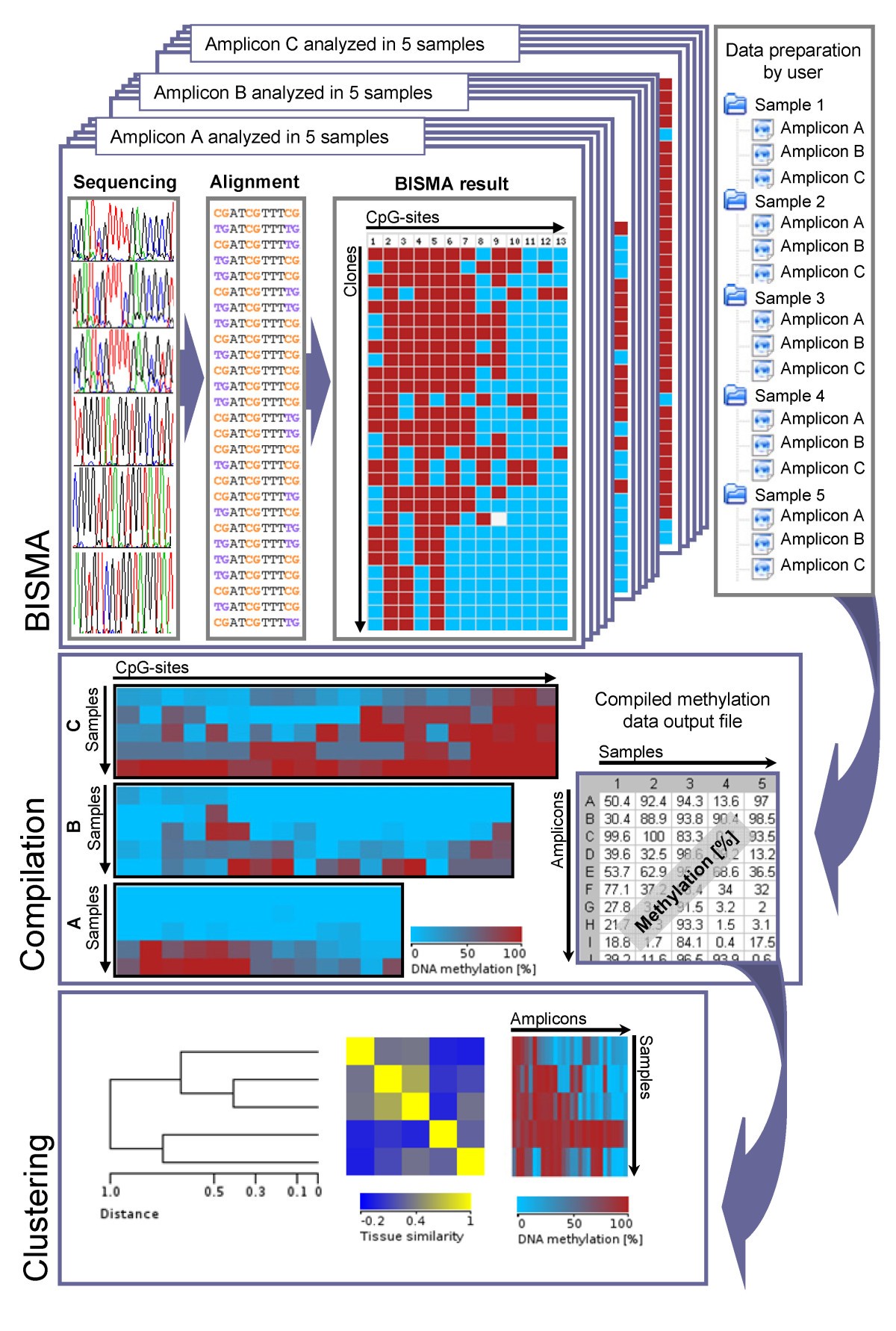
BISMA - Fast and accurate bisulfite sequencing data analysis of individual clones from unique and repetitive sequences | BMC Bioinformatics | Full Text
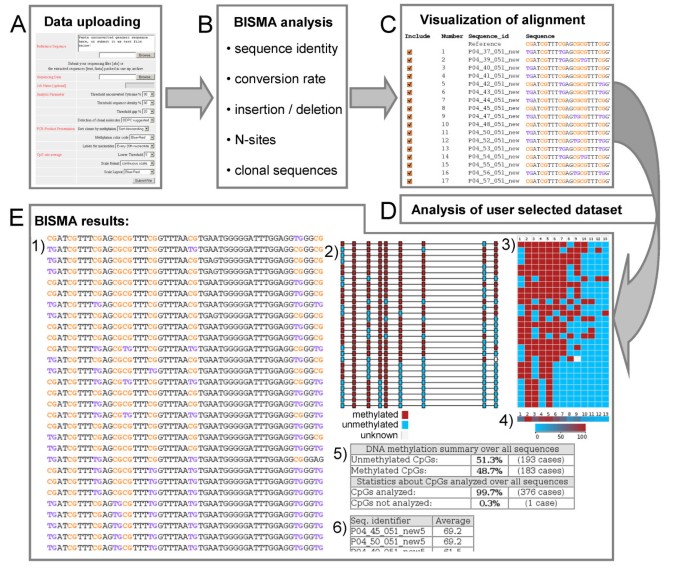
BISMA - Fast and accurate bisulfite sequencing data analysis of individual clones from unique and repetitive sequences | BMC Bioinformatics | Full Text

Bisulfite sequencing validation of MBD-Seq result (e.g., MBD22 region) a) Methylation concentration levels from MBD-seq b) Bisulfite sequencing results.
Methy-Pipe: An Integrated Bioinformatics Pipeline for Whole Genome Bisulfite Sequencing Data Analysis | PLOS ONE
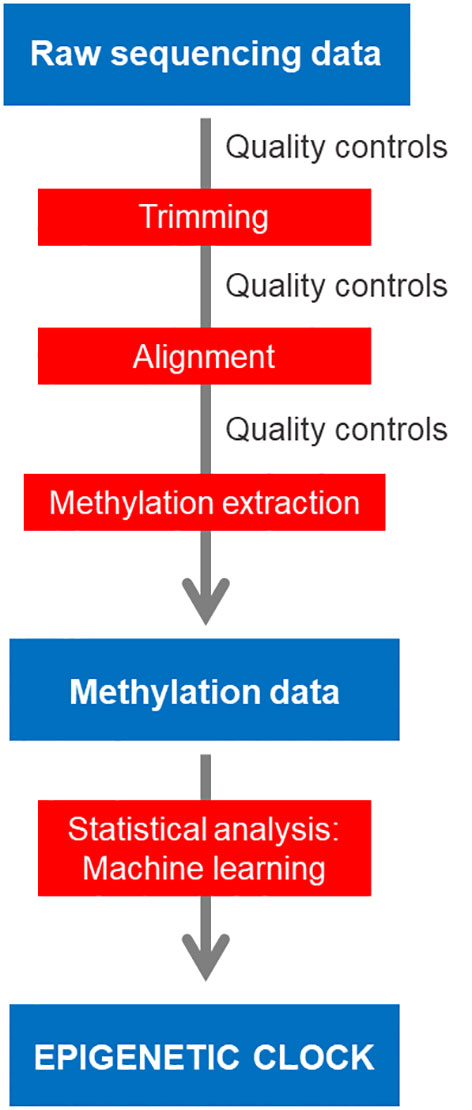
Frontiers | Bioinformatic analysis for age prediction using epigenetic clocks: Application to fisheries management and conservation biology