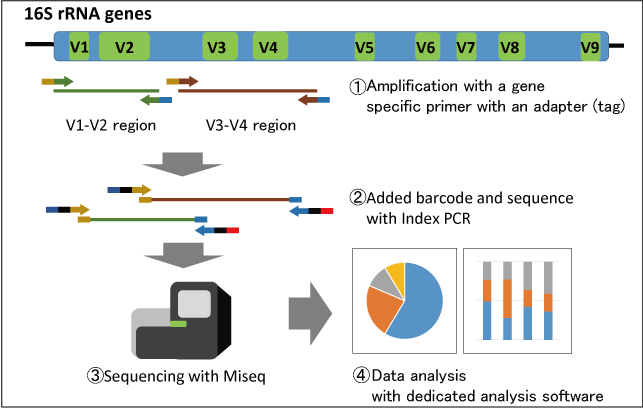
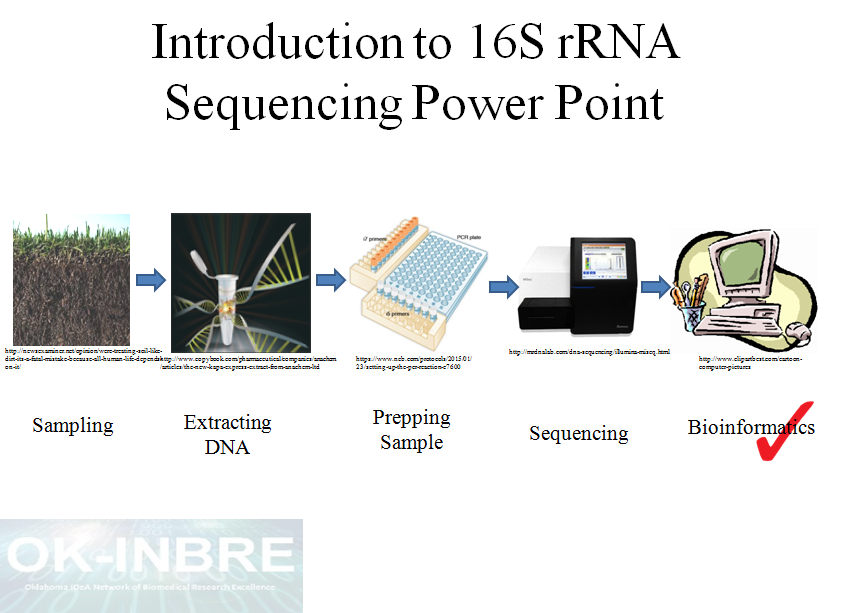
Research Techniques Made Simple: Bacterial 16S Ribosomal RNA Gene Sequencing in Cutaneous Research. - Abstract - Europe PMC

Multicenter assessment of microbial community profiling using 16S rRNA gene sequencing and shotgun metagenomic sequencing - ScienceDirect

Research Techniques Made Simple: Bacterial 16S Ribosomal RNA Gene Sequencing in Cutaneous Research - ScienceDirect
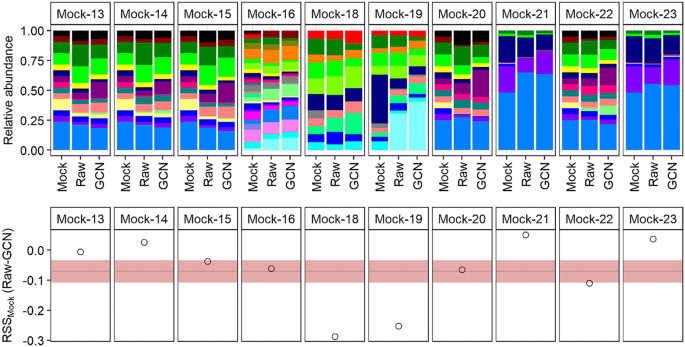
16S rRNA Gene Copy Number Normalization Does Not Provide More Reliable Conclusions in Metataxonomic Surveys | SpringerLink
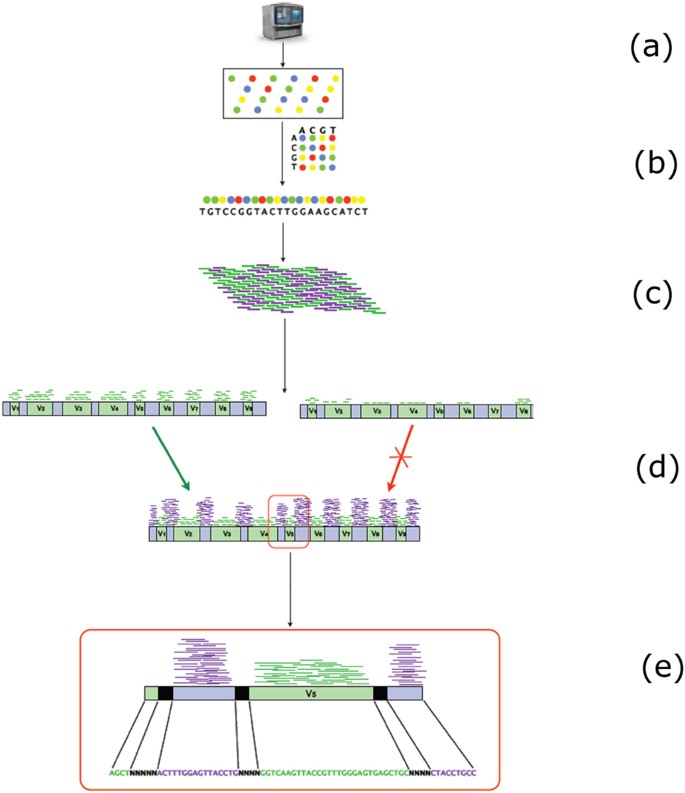
Direct 16S rRNA-seq from bacterial communities: a PCR-independent approach to simultaneously assess microbial diversity and functional activity potential of each taxon | Scientific Reports
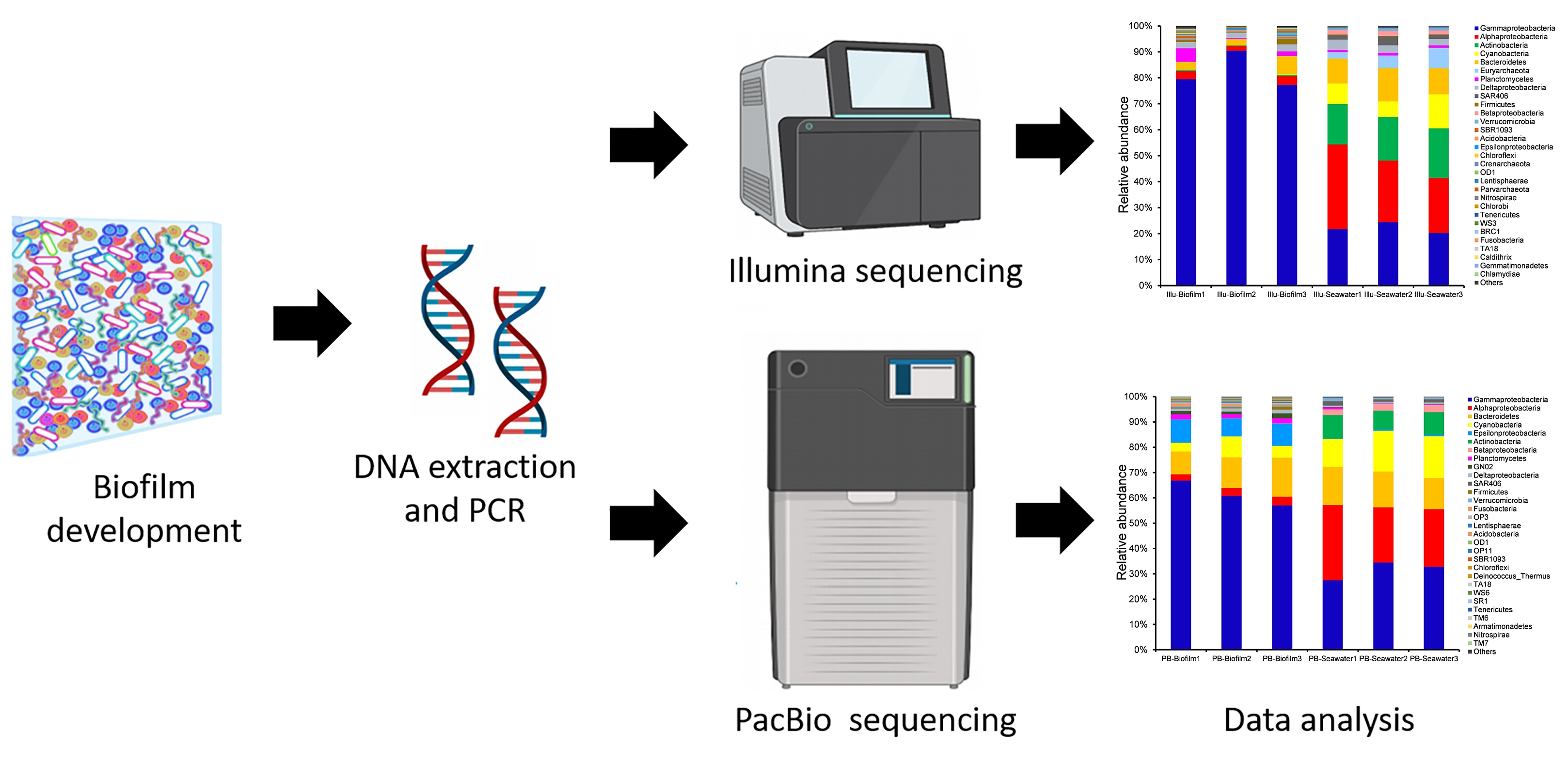
Genes | Free Full-Text | Microbial Richness of Marine Biofilms Revealed by Sequencing Full-Length 16S rRNA Genes
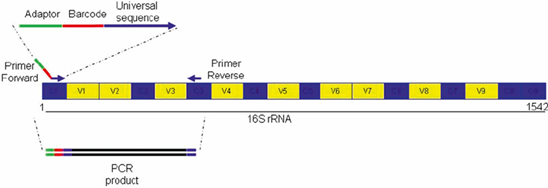
16s rRNA Sequencing Analysis | Microbiome Sequencing – 1010Genome | Quality NGS Bioinformatics Data Analysis Services
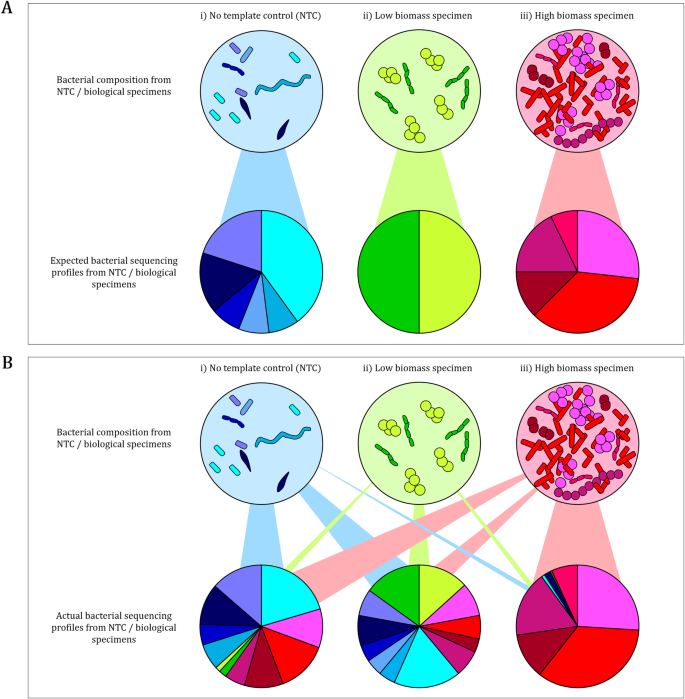
Optimizing 16S rRNA gene profile analysis from low biomass nasopharyngeal and induced sputum specimens | BMC Microbiology | Full Text
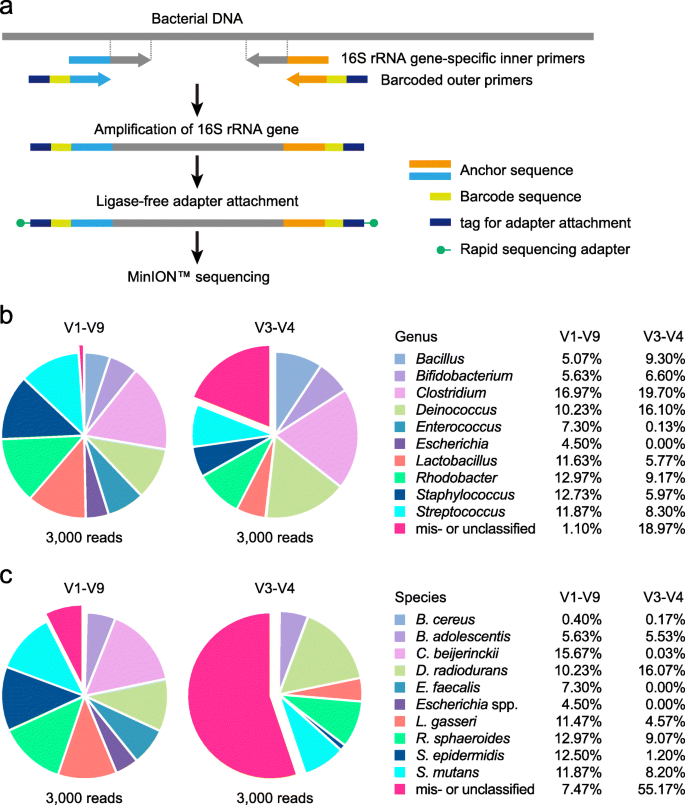
Full-length 16S rRNA gene amplicon analysis of human gut microbiota using MinION™ nanopore sequencing confers species-level resolution | BMC Microbiology | Full Text
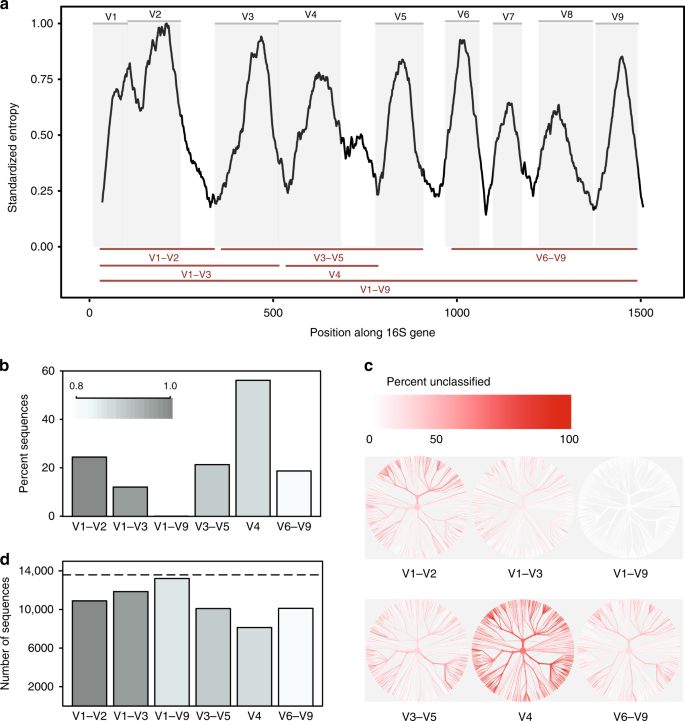
Evaluation of 16S rRNA gene sequencing for species and strain-level microbiome analysis | Nature Communications