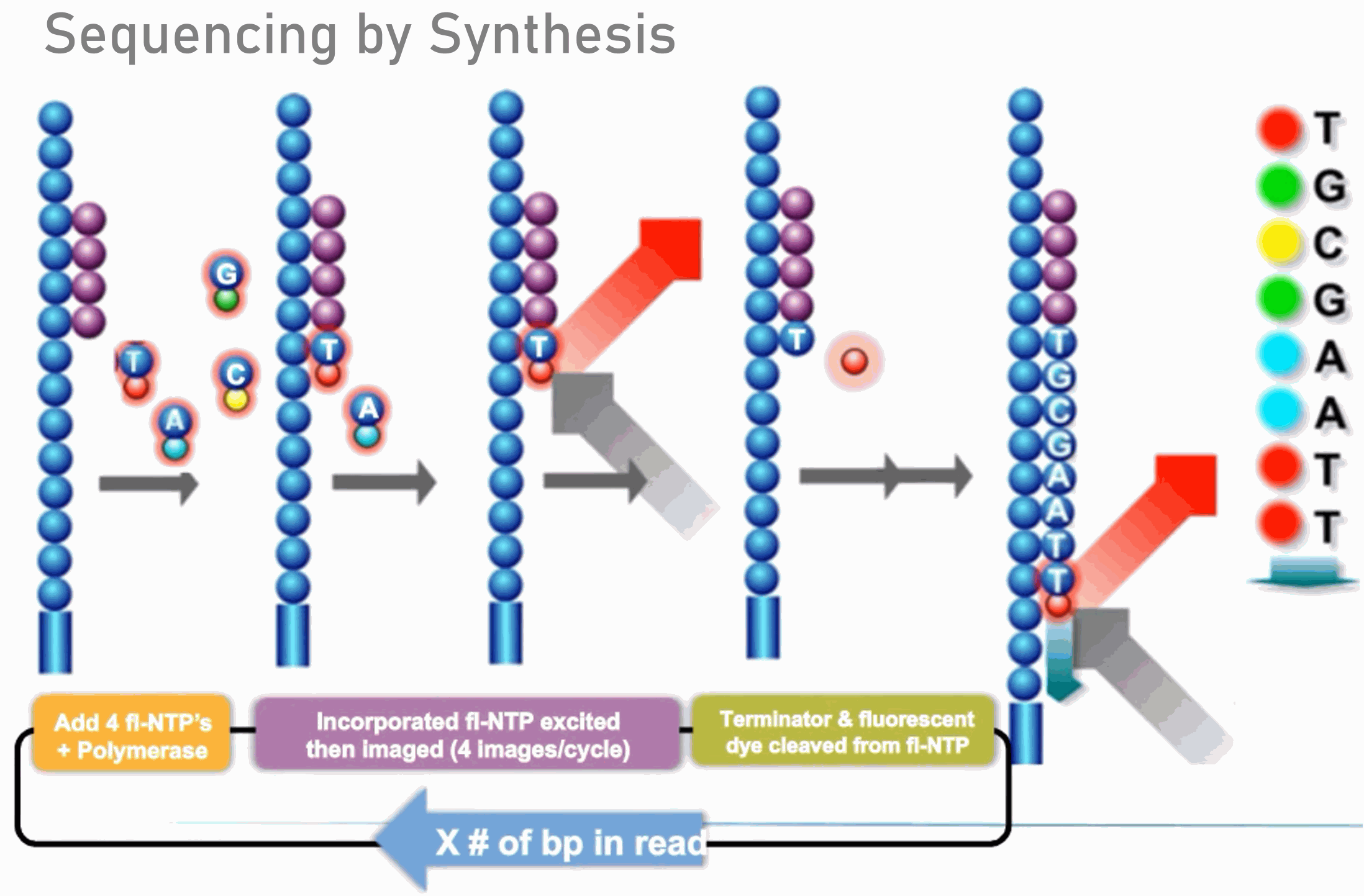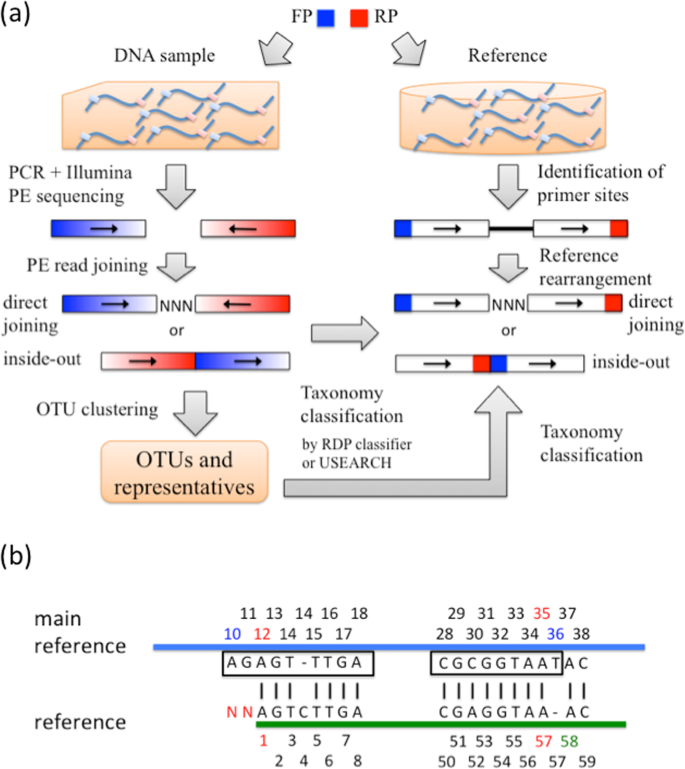
Joining Illumina paired-end reads for classifying phylogenetic marker sequences | BMC Bioinformatics | Full Text

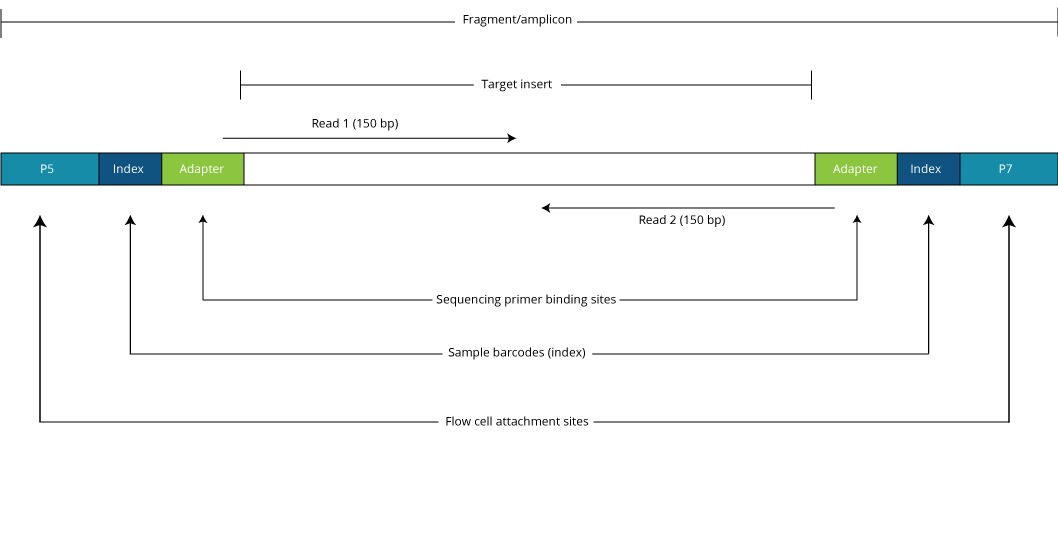
Scheme of the MiSeq sequencing library in this study. The 150-bp paired... | Download Scientific Diagram
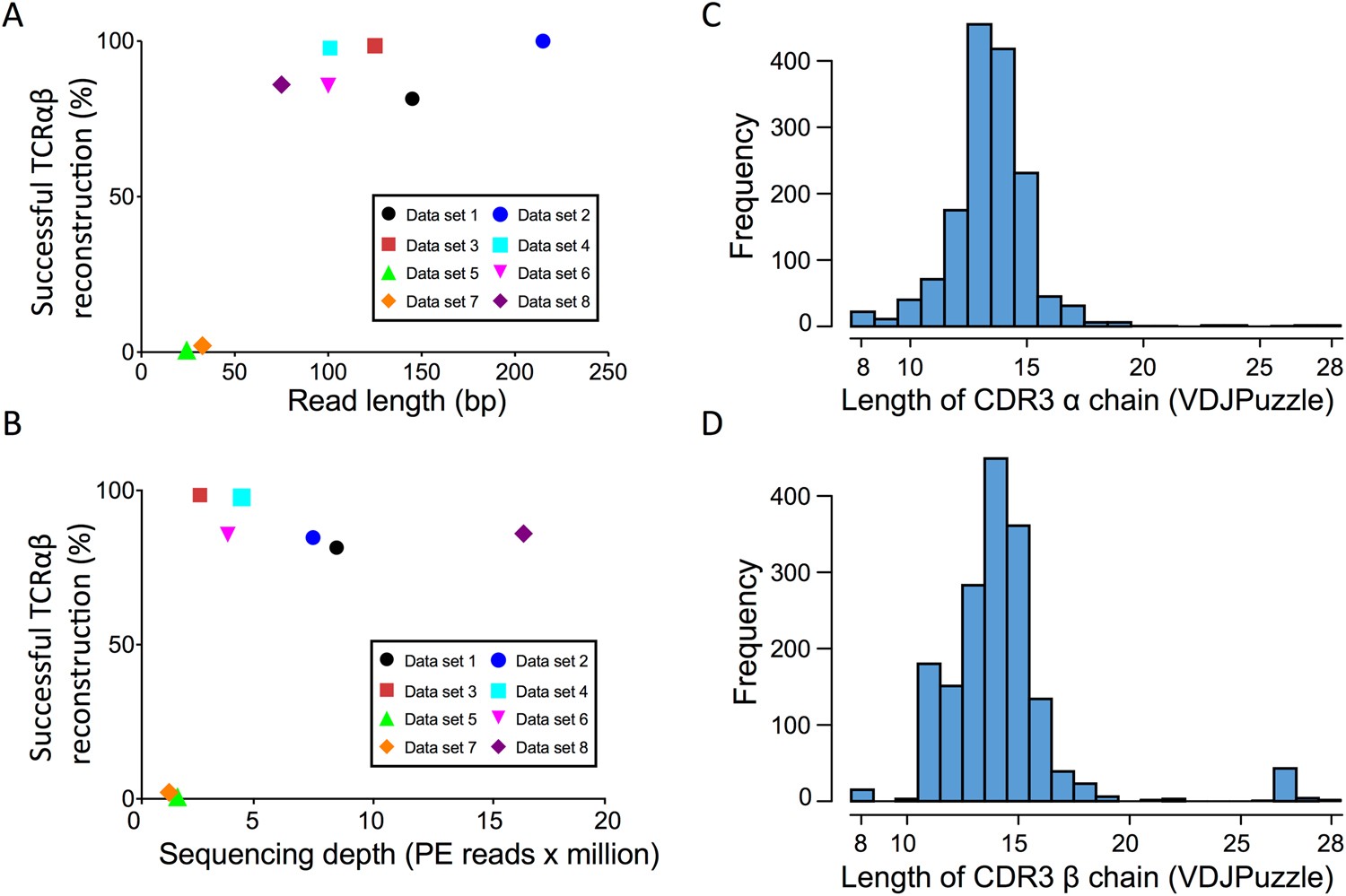
Impact of sequencing depth and read length on single cell RNA sequencing data of T cells | Scientific Reports

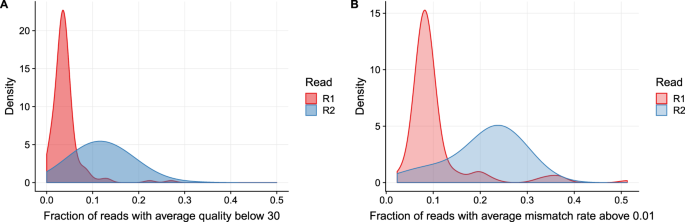
Quality assessment of raw FASTQ sequence data for 150 bp paired end... | Download Scientific Diagram

A comprehensive evaluation of single-end sequencing data analyses for environmental microbiome research | SpringerLink
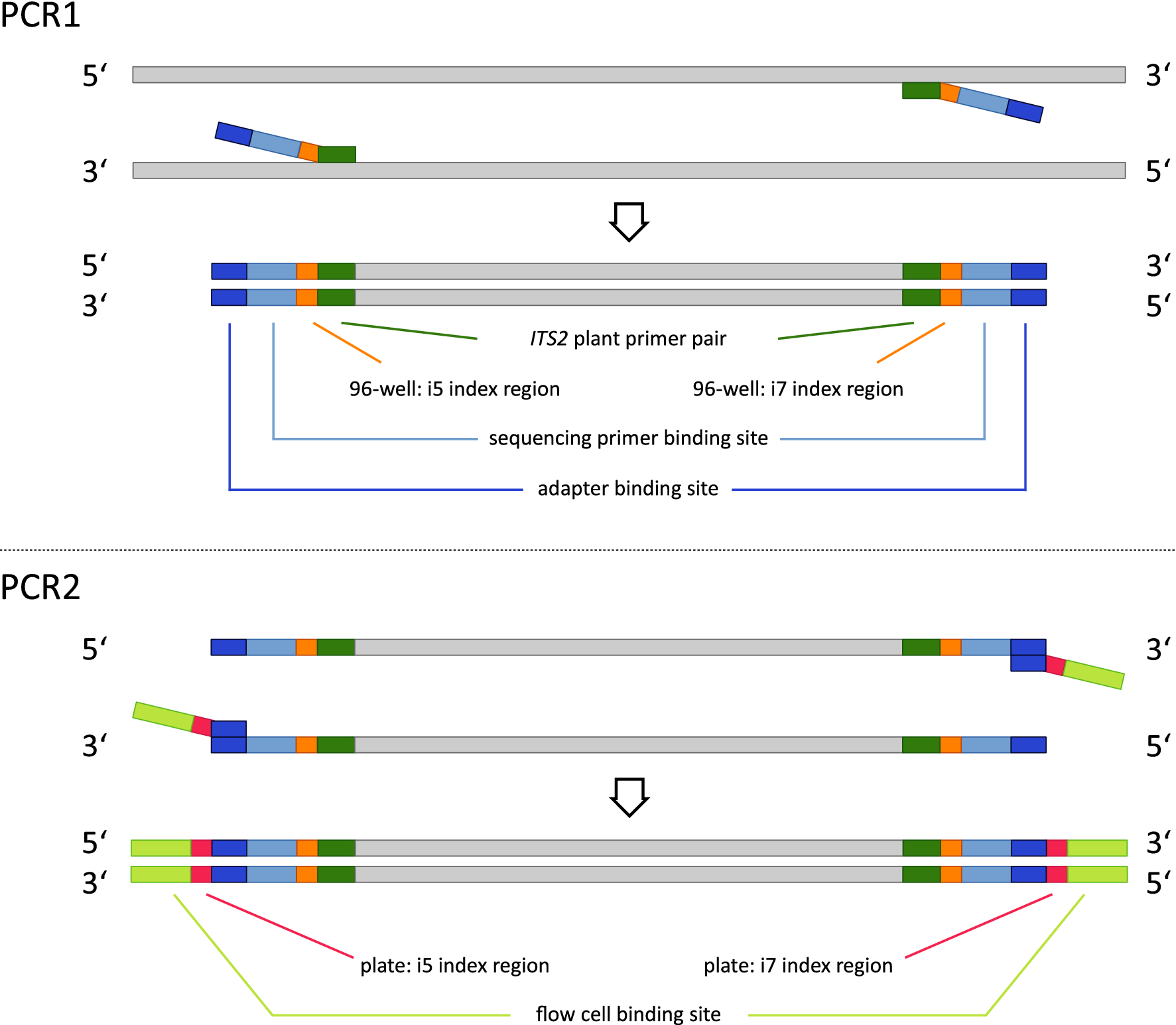
Handling of targeted amplicon sequencing data focusing on index hopping and demultiplexing using a nested metabarcoding approach in ecology | Scientific Reports
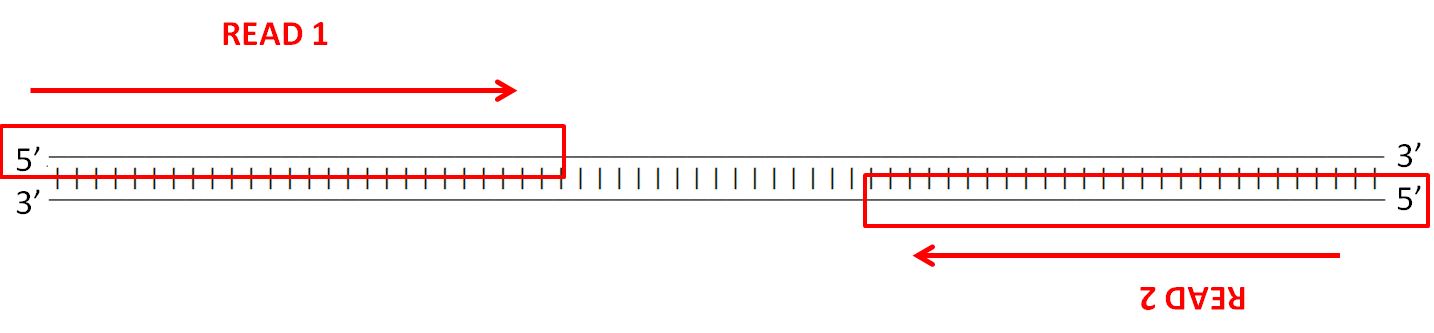
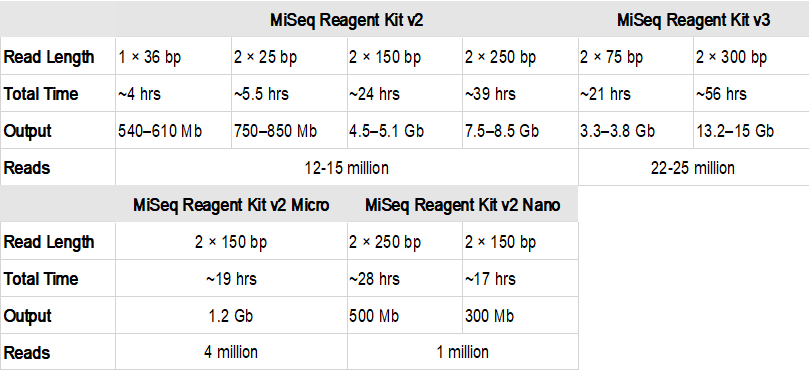
What 2 indicates in 2*150 bp, 2*300 bp maximum read length from Illumina website http://www.illumina.com/systems/sequencing-platforms.html?

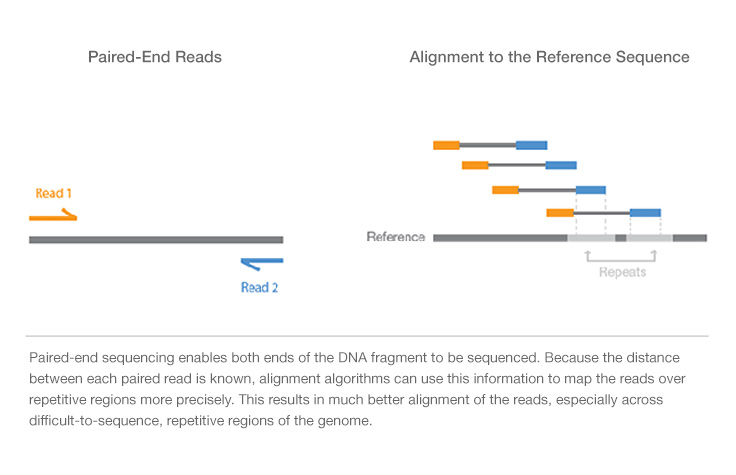
Paired-end sequencing (left) showing Read 1 and Read 2 primers starting... | Download Scientific Diagram